Credit: Nicholas Stephanopoulos
The central goal of nanotechnology is the manipulation of materials on an atomic or molecular scale, especially to build microscopic devices or structures. Three-dimensional cages are one of the most important targets, both for their simplicity and their application as drug carriers for medicine. DNA nanotechnology uses DNA molecules as programmable “Legos” to assemble structures with a control not possible with other molecules.
However, the structure of DNA is very simple and lacks the diversity of proteins that make up most natural cages, like viruses. Unfortunately, it is very difficult to control the assembly of proteins with the precision of DNA. That is, until recently. Nicholas Stephanopoulos — an assistant professor in Arizona State University’s Biodesign Center for Molecular Design and Biomimetics, and the School of Molecular Sciences — and his team built a cage constructed from both protein and DNA building blocks through the use of covalent protein-DNA conjugates.
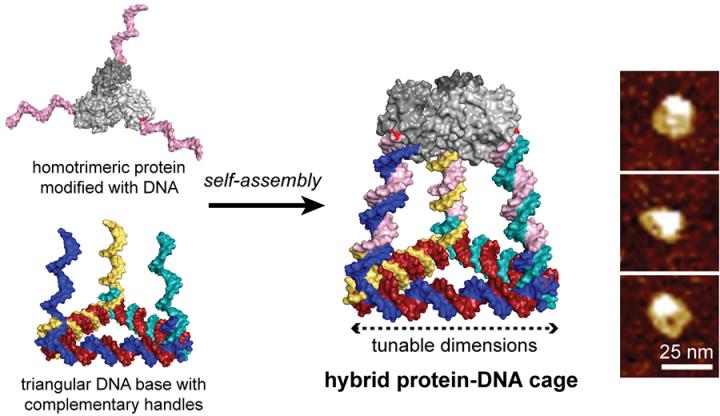
In a paper published in ACS Nano, Stephanopoulos modified a homotrimeric protein (a natural enzyme called KDPG aldolase) with three identical single strand DNA handles by functionalizing a reactive cysteine residue they introduced onto the protein surface. This protein-DNA “Lego” was co-assembled with a triangular DNA structure bearing three complementary arms to the handles, resulting in tetrahedral cages comprised of six DNA sides capped by the protein trimer. The dimensions of the cage could be tuned through the number of turns per DNA arm and the hybrid structures were purified and characterized to confirm the three-dimensional structure.
Cages were also modified with DNA using click chemistry, which is a customized type of chemistry, to create elements rapidly with great reliability joining microscopic units together demonstrating the generality of the method.
“My lab’s approach will allow for the construction of nanomaterials that possess the advantages of both protein and DNA nanotechnology, and find applications in fields such as targeted delivery, structural biology, biomedicine, and catalytic materials,” Stephanopoulos said.
Stephanopoulos and his team see an opportunity with hybrid cages — merging self-assembling protein building blocks with a synthetic DNA scaffold — that could combine the bioactivity and chemical diversity of the former with the programmability of the latter. And that is what they set out to create — a hybrid structure constructed through chemical conjugation of oligonucleotide (a synthetic DNA strand) handles on a protein building block. The triangular base bearing three complementary single-stranded DNA handles is self-assembled and purified separately by heating it to alter its properties.
“We reasoned that by designing these two purified building blocks, they would spontaneously snap together in a programmable way, using the recognition properties of the DNA handles,” Stephanopoulos said. “It was especially critical to use a highly thermally stable protein like this aldolase, because this self-assembly only works at 55 degrees Celsius, and many proteins fall apart at those temperatures.”
Another advantage of DNA, which is not possible with proteins, is tuning the cage size without having to redesign all the components. Stephanopoulos continued, “The size of this assembly could then be rationally tuned by changing the length of each DNA edge, whereas the protein would provide a scaffold for the attachment of small molecules, targeting peptides or even fusion proteins.”
While other examples of hybrid structures exist, this particular cage is the first one constructed through chemical conjugation of oligonucleotide handles on a protein building block. This strategy can in principle be expanded to a wide range of proteins (some with cancer targeting abilities, for example). Thus, Stephanopoulos’s work has the potential to enable a whole new hybrid field of protein-DNA nanotechnology with applications not possible with either proteins or DNA alone.
###
Media Contact
Kimberley Baptista
[email protected]
Original Source
https:/
Related Journal Article
http://dx.




