New findings reveal many different structural models, which can eventually lead to developing more targeted antiviral vaccines.
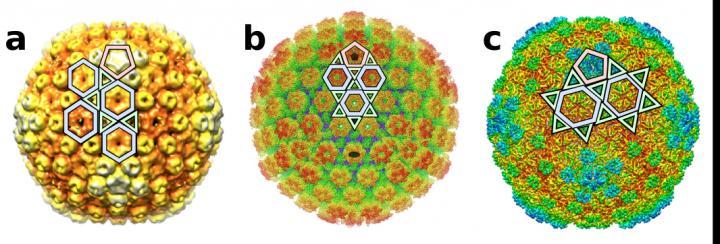
Credit: Antoni Luque, San Diego State University and Reidun Twarock, University of York.
New research reveals that the way viruses were perceived in terms of their architecture will need to be retooled, because they are actually structured in many more patterns than previously understood. The findings could have significant impact on how they are classified, our understanding of how they form, evolve and infect hosts, and strategies to identify ways to design vaccines to target them.
In the 1950s and ’60s as scientists began to obtain high resolution images of viruses, they discovered the detailed structure of the capsid – an outer protective layer composed of multiple copies of the same protein – which protects the virus’ genetic material. The majority of viruses have capsids that are typically quasi-spherical and display icosahedral symmetry – like a 20-sided dice for instance.
The capsid shell is what protects them, and as scientists discovered their structure, they proposed that capsids could have different sizes and hold different amounts of genome, and therefore could infect hosts differently.
Why this matters
When designing drugs to target viruses, scientists can now take their varying structural shapes into account to improve efficacy.
Two researchers who study the structures of viruses, Antoni Luque, a theoretical biophysicist at San Diego State University and a member of its Viral Information Institute, and Reidun Twarock, a mathematical biologist from the University of York, UK, and a member of York’s Cross-disciplinary Centre for Systems Analysis, show that many viruses have essentially been misclassified for 60 years, including common viruses such as Herpes simplex and Zika.
This was because despite having the structural images from cryo-electron microscopy, we did not have the mathematical description of many of the architectures of different viruses.
“We discovered six new ways in which proteins can organize to form icosahedral capsid shells,” Luque said. “So, many viruses don’t adopt only the two broadly understood capsid architectures. There are now at least eight ways in which their icosahedral capsids could be designed.”
They used a generalization of the quasiequivalence principle to see how proteins can wrap around an icosahedral capsid.
Their study, which will be published in Nature Communications on Friday, September 27, also shows that viruses that are part of the same structural lineage, based on the protein that they’re composed of, adopt consistent icosahedral capsid layouts, providing a new approach to study virus evolution.
Biotech applications
Structural biologists can now take this information and reclassify the structure of the viruses, which will help unveil molecular and evolutionary relationships between different viruses.
It will also provide a guide to engineer new molecular containers for nanotech and biotech applications, and it will help scientists to identify specific strategies to target the assembly of proteins in the capsid. This can eventually lead to a more systematic approach to developing antiviral vaccines.
“We can use this discovery to target both the assembly and stability of the capsid, to either prevent the formation of the virus when it infects the host cell, or break it apart after it’s formed,” Luque said. “This could facilitate the characterization and identification of antiviral targets for viruses sharing the same icosahedral layout.”
This new framework accommodates viruses that were previously outliers, provides new predictions of viral capsid architectures, and has identified common geometrical patterns among distant evolutionary related viruses that infect everyone from humans to bacteria.
Twarock said the new blueprints also provide “a new perspective on viral evolution, suggesting novel routes in which larger and more complex viruses may have evolved from simple ones at evolutionary timescales.”
Architectural applications
The geometries could be also used in new architectural designs in buildings and construction.
Since the 1960s, these viral capsids have been classified using the geometrical framework introduced by structural biologist Donald Caspar and biophysicist Aaron Klug, which were inspired by the geodesic domes designed by the renowned architect R. Buckminster Fuller. However, as molecular imaging techniques have advanced, an increasing number of 3D viral capsid reconstructions that included viruses like Herpes or Zika have fallen out from this classical geometrical framework.
“This study introduces a more general framework than the classic Caspar-Klug construction. It is based on the conservation of the local vertices formed by the proteins that interact in the capsid,” Luque explained. “This approach led to the discovery of six new types of icosahedral capsid layouts, while recovering the two classical layouts from Caspar-Klug based on Goldberg and geodesic polyhedra.”
###
Collaborations and funding
Co-authors Antoni Luque from San Diego State University and Reidun Twarock from University of York began collaborating on this research in 2017. Luque’s lab studies the architecture and ecology of viruses using mathematical and computational models. Using this new framework from this research project, he is currently developing methods to investigate the architecture of viruses in different environments, which could have implications in medicine, ecology, and evolution.
Twarock has been developing geometric models for virus architecture since 2004. Her group is developing mathematical and computational approaches to investigate the consequences of viral geometry for mechanisms in viral life cycles and viral evolution.
Twarock was funded by the EPSRC (Engineering and Physical Sciences Research Council), the Royal Society, and the Wellcome Trust.
This paper is embargoed until 5 a.m EST on Friday, September 27th, 2019 when you can access it at:
https:/
Media Contact
Padma Nagappan
[email protected]




