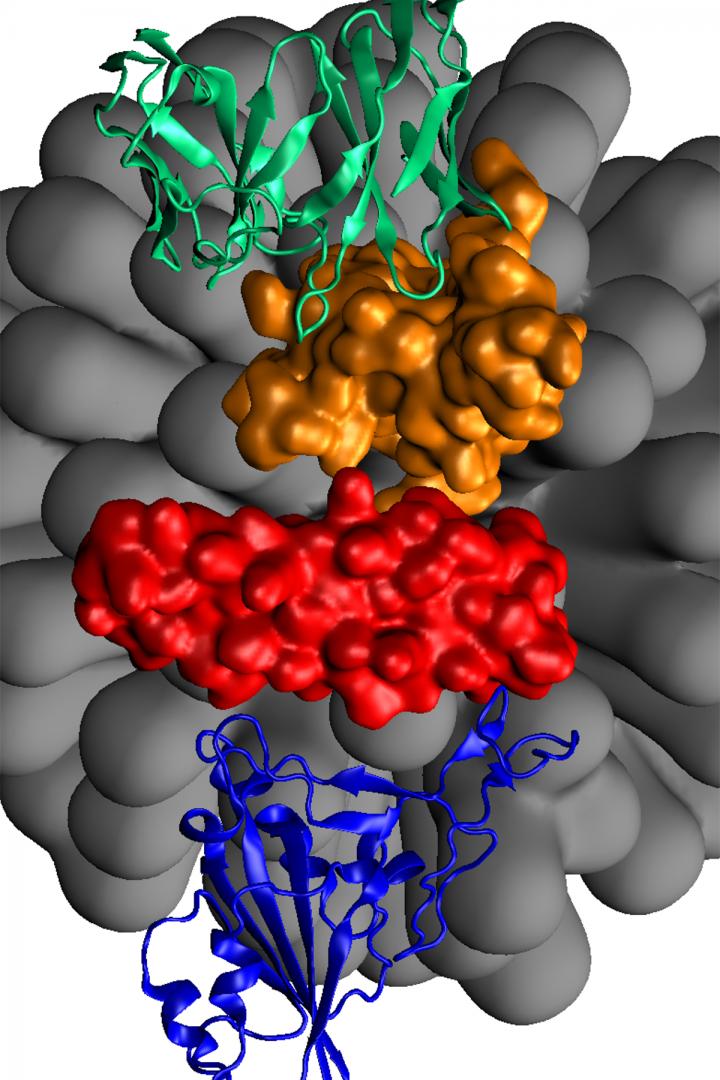
Credit: UIC
The SARS-CoV-2, the new coronavirus behind the current pandemic, infects humans by binding its surface-exposed spike proteins to ACE2 receptors exposed on the cell membranes.
Upon a vaccination or a real infection, it takes several weeks before the immunity develops antibodies that can selectively bind to these spike proteins. Such antibody-labeled viruses are neutralized by the natural killer and T cells operated by the human immunity.
An alternative approach to train the immunity response is offered by researchers at the University of Illinois Chicago and California State University at Sacramento who have developed a novel strategy that redirects antibodies for other diseases existing in humans to the spike proteins of SARS-CoV-2.
In their study published by the Journal of Physical Chemistry Letters, the team proposes using peptide-based double-faced “booster” inhibitors, with one face binding to the spike proteins of SARS-CoV-2 and the other face binding to generic hepatitis B antibodies.
“Once the SARS-CoV-2 viruses become labeled by the hepatitis B antibodies via intermediate boosters, the viruses will be neutralized. This universal approach allows a dramatic shortening of the response time upon real infections, which can be critical in certain patients or conditions,” said Petr Král, UIC professor of chemistry, physics, pharmaceutical sciences and chemical engineering, and senior author on the paper.
Král and Yanxiao Han, who recently earned a Ph.D. in chemistry at UIC and is first author on the paper, believe the study could provide guidance in the preparation of generic therapeutics against emerging pathogens with the combined advantages of small-protein and antibody therapies.
“The dramatic impact which novel viruses can have on humans could be fast mitigated in the absence of their vaccination if generic antibodies present within them are temporarily retrained to recognize these viruses,” the researchers wrote.
In a study published last spring, Král and Han extracted different peptides from ACE2 that interact directly with the viral spike protein.
“We investigated potential COVID-19 therapeutics using computer simulations based on the X-ray crystal structure of the receptor-binding domain of SARS-CoV-2 when it is bound to ACE2,” Král said. “Similar to our latest study, identifying these kinds of inhibitors could lead to new treatments to combat the coronavirus.”
###
Co-author on the paper is Katherine McReynolds of California State University at Sacramento.
This work is supported by funding from the UIC Center for Clinical and Translational Science.
Media Contact
Brian Flood
[email protected]
Original Source
https:/
Related Journal Article
http://dx.



