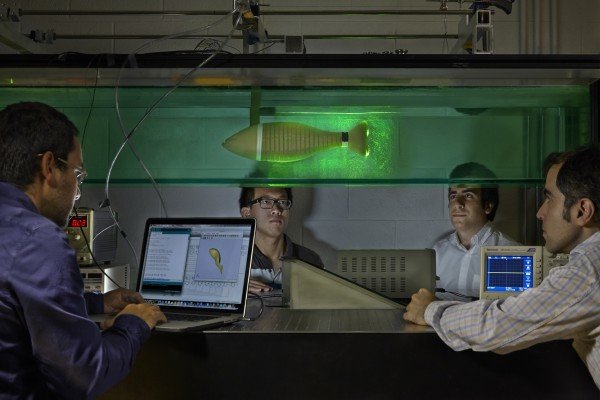
Embracing the idea that molecules can be programmed much like a computer, researchers can now perform remarkable feats on a very small scale. New Caltech faculty member Lulu Qian, assistant professor of bioengineering, performs research in the field of molecular programming because it allows her to design synthetic molecular systems with neural-network-like behaviors and tiny robots, both from the programmed interactions of DNA molecules.

Originally from Nanjing, China, Qian received her bachelor’s degree from Southeast University in 2002 and her PhD from Shanghai Jiao Tong University in 2007. After working as a postdoctoral scholar at Caltech in the laboratory of Shuki Bruck, Qian became a visiting fellow at Harvard University; she returned to Caltech and joined the faculty in July. Recently, Qian answered a few questions about her research, and how it feels to be back at Caltech.
What do you work on?
I work on rationally designing and creating molecular systems with programmable behaviors. I am interested in programming biological molecules—like DNA and RNA—to recognize molecular events from the biochemical environment, process information, make decisions, take actions, and to learn and evolve. Molecular programming is not just about using computer programs to aid the design and analysis of molecular systems; it is more about adapting the principles of computer science to create biochemical systems that can carry out instructions to perform tasks at the molecular level. For example, I develop simple and standard molecular components that can be used to perform a variety of tasks and systematic ways to configure the behavior of interacting molecules to carry out one computational or mechanical task or another. These custom-designed molecules can be ordered from a commercial supplier and mixed in a test tube to generate a “molecular program.”
Using this approach, I have designed DNA circuits that can solve basic logic problems, and I have constructed a DNA neural network that can perform simple associative memory functions—much like a network of neurons in the brain, though in a rudimentary way. In my future research, I would like to improve the speed, robustness, and complexity of these implementations and to explore the possibility of creating molecular systems with learning capabilities, while also beginning new work in the field of molecular robots—tiny, nanoscale machines made of DNA that can perform a designed task such as sorting cargo or solving a maze.
What do you find most exciting about your research?
I am driven by curiosity—outside of the lab, I like Legos and puzzles—and I view life as a program, one that is much more sophisticated than any other program that we know of so far. The sequence of nucleotides that make up DNA—As, Ts, Cs, and Gs—encodes the program within a genome, orchestrating molecules to sense, to compute, to respond, and to grow. Because of their different lengths and sequences, one genome produces a bacterium while another produces a plant, or an insect, or a mammal. The genetic program describes how to make molecules, and molecules are machines that can achieve complex tasks to regulate the behaviors of individual cells. To better appreciate the molecular programs that nature creates, I want to understand what possible behaviors a network of interacting molecules can exhibit and how we can rationally design such behaviors.
But, I am also driven by my engineering nature. I want to design and build molecular systems with increasing complexity and sophistication. For example, you could imagine using such molecular machines to make a nanoscale factory that manufactures novel chemicals in a test tube. These chemicals could become new materials or new drugs. You could also imagine embedding such molecular machines into individual cells so that you could collect information from the molecular environment and regulate the cell’s behavior. Such regulation could lead to responsive biosynthesis—the production of proteins or other molecules in response to a stimulus—or localized diagnostics followed by therapeutics.
How did you get into your field?
I started programming computers when I was 13 years old, and I have loved it ever since then. My dad was a philosopher, and because of his influence, I got curious about fundamental questions such as who I am and why I think the way that I do. At first, I tried to look for these answers in molecular biology, but as a programmer, biology was difficult for me to understand. Unlike in programming, you cannot just define a few logical principles to understand the behavior of an entire biological system or organism. At the time, biology was not as fun for me—or as logical—as computer programming.
But just before I went to graduate school, I discovered the first publication in DNA computing by Len Adleman at the University of Southern California. He used DNA molecules as a computing substrate to solve a hard math problem. The moment that I finished reading this paper, I felt completely excited. It was the first time that I saw a strong relationship between molecules that are traditionally only used in biology—like DNA and RNA—and computer programming. That was when I started working in my field.
Story Source:
The above story is based on materials provided by California Institute of Technology, Jessica Stoller-Conrad.