New methods for understanding the immune system and how to boost it
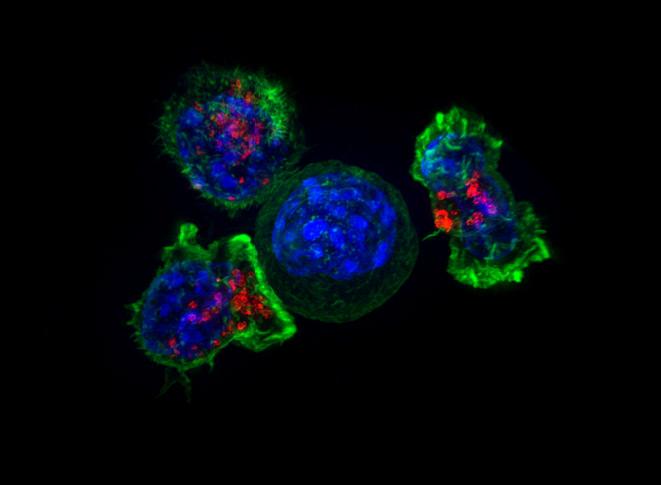
Credit: National Institutes of Health
The human body’s immune system is like a vast team of special agents. Certain cells called T cells each individually specialize in recognizing a particular intruder, such as the influenza virus or salmonella. Determining a given T cell’s target is a critical step in designing personalized treatments for cancers and developing vaccines. Now, a team of Caltech scientists has developed two new methods for rapidly determining T cell targets.
The work was done in the laboratory of David Baltimore, Robert Andrews Millikan Professor of Biology and president emeritus. Two papers describing the research appear in the January 28 issue of the journal Nature Methods.
A cell that is infected with a pathogen–for example, an influenza virus–will display bits of the invader’s genetic material on the cell surface, like waving a red flag to indicate what is going on inside the cell. These “flags,” called antigens, are presented on proteins on the cell surface, called MHCs (major histocompatibility complexes). Each T cell is specialized to recognize a different antigen. When a T cell finds a cell displaying its target antigen on an MHC complex, the T cell will bind to it and destroy it.
There are from 1 million to 5 million unique T cells on average in a human targeting countless different pathogens. Though scientists can characterize the function and molecular makeup of a T cell’s receptor, it is difficult to determine what target a given receptor specifically recognizes. In fact, fewer than 1,000 antigen-T cell pairs are known.
Now, led by postdoctoral scholars Alok Joglekar and Guideng Li, researchers in the Baltimore laboratory have developed two new methods for determining the targets of T cells.
In the first method, the scientists attached proteins, called signaling domains, onto MHCs. The new complex, called a signaling and antigen-presenting bifunctional receptor, or SABR, is designed to send a signal into the cell to make it glow bright green once it has been bound by a corresponding T cell. A researcher could then take thousands of different antigens, each presented by a SABR, and combine them with a particular T cell. Only the cells presenting the correct antigen should glow green, allowing the researchers to fish out the correct antigen–the T cell’s target.
The second method takes advantage of a natural phenomenon called trogocytosis. This occurs when a T cell and its target cell bind together and exchange proteins that are bound to their surfaces. Although researchers have not yet determined why trogocytosis occurs, the Baltimore laboratory decided to use the phenomenon to indicate T-cell targets. To do this, the researchers made a pool of antigen-presenting cells, each displaying a unique antigen, and then exposed them to T cells with a receptor of interest. Only the cells presenting the correct antigen acquired markers from the T cell via trogocytosis. Afterward, the antigen corresponding to the T cell could then be identified by the marker on its surface.
Understanding antigen-T cell pairs is crucial for developing cancer vaccines and also for designing personalized treatments for cancers, as antigens can also be signatures of cancer. No two cancers are the same, and because cancer cells grow so rapidly, they also mutate rapidly. Ideally, then, a scientist could take a sample of a person’s tumors, isolate T cells from them, and use one of these methods to discover the antigens that are targeted by the T cells. Once these targets are identified, the T cells can be used to augment the patient’s own immune system in various ways to help it better fight the person’s cancer.
###
A paper describing the SABR method is titled “T cell antigen discovery via Signaling and Antigen presenting Bifunctional Receptors.” Postdoctoral scholar Alok Joglekar is the study’s first author. In addition to Joglekar and Baltimore, co-authors are Caltech research technicians Michael Leonard and Margaret Swift; former Caltech research technician John Jeppson; postdoctoral scholar Guideng Li; former Caltech undergraduate and current research technician Stephanie Wong (BS ’16); former Caltech postdoctoral scholar Songming Peng now of PACT Pharma; Jesse Zaretsky of UCLA; James Heath, a former Caltech professor now at the Institute for Systems Biology in Seattle; Antoni Ribas of UCLA; and former postdoctoral scholar Michael Bethune.
A paper describing the trogocytosis method is titled “T cell antigen discovery via trogocytosis.” Guideng Li and Michael Bethune are the study’s first authors. In addition to Li, Bethune, and Baltimore, co-authors are Stephanie Wong; Alok Joglekar; Michael Leonard; undergraduate Jessica Wang; former graduate student Jocelyn Kim (PhD ’16), now of UCLA; Donghui Cheng of UCLA; Songming Peng; Jesse Zaretsky; Caltech graduate students Yapeng Su and Yicheng Luo; James Heath; Antoni Ribas; and Owen Witte of UCLA. Baltimore, Ribas, Heath, and Witte are also members of the Parker Institute for Cancer Immunotherapy at UCLA and Caltech.
Media Contact
Lori Dajose
[email protected]
626-658-0109
Original Source
http://www.
Related Journal Article
http://dx.