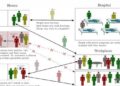
The discovery by UMD researchers suggests ‘functional mosaicism’ could explain some antibiotic resistance and cancer relapse
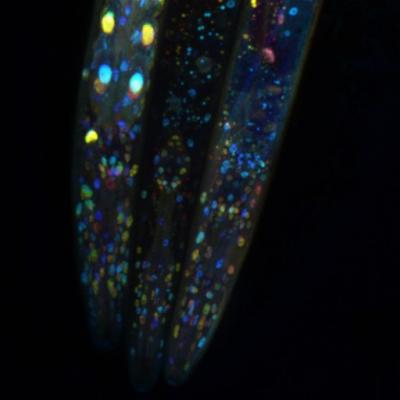
Credit: Snusha Ravikumar/University of Maryland
University of Maryland researchers have discovered that seemingly identical cells can use different protein molecules to carry out the same function in an important cellular process. The scientists named this newly discovered variability “functional mosaicism,” and it has significant implications for the development of therapeutic treatments, which are often designed to target a specific molecule, or a gene that produces a specific molecule.
By showing that individual cells within the same tissue can use different molecules to carry out the same functions, the new findings may help explain the persistence of some infected cells after drug treatment and relapses in complex diseases such as cancer. The research paper describing the work was published in the September 10, 2019, issue of the journal Nucleic Acids Research.
“Functional mosaicism introduces a cautionary message,” said Antony Jose, senior author of the paper and associate professor of cell biology and molecular genetics at UMD. “It means that if you’re developing a drug therapy that targets a certain process within cells, you can no longer assume that all cells of the same type use the same molecule or molecules to carry out that process. If you do, you may miss a few cells where things work differently.”
Although scientists have long known about biological redundancy–when cells can carry out important functions in multiple ways–they typically assume consistency and the same redundancies among all cells of the same type. Functional mosaicism, on the other hand, means that within a group of seemingly identical cells, some of the individual cells can use an alternate path for certain functions.
“With biological redundancy, if you’re developing a drug to target a particular function and the cells have an alternative way to accomplish the function, your drug will fail because all cells will use the alternative,” Jose explained. “Functional mosaicism is different, and it can fool you. Your drug may appear to work because it works in some cells – even most cells, but there may be a few cells that persist because they can use an alternative you’re not aware of.”
Jose and his team of UMD researchers discovered functional mosaicism while studying RNA interference (RNAi), an important biological process that helps cells fight infection. Although they have not looked for functional mosaicism in other cellular processes yet, their findings represent a previously unknown and unexpected level of variability that, according to Jose, is likely to exist in many other cellular processes.
In the study, Jose and his team were investigating how RNA moves from one tissue type to another to turn off genes. Like DNA, RNA is a string of coded instructions that guide various functions in living cells. Unlike DNA, RNA usually occurs in single strands, except in some viruses, which introduce double strands of RNA into cells. One common defense that living cells have developed against viral infection is that they recognize double-stranded RNA and turn off genes with matching DNA sequences. This gene-silencing process is called RNAi.
RNAi relies on a group of protein molecules, or enzymes, called RNA-dependent RNA polymerases (RdRp), which replicate strands of RNA that can find and silence their matching genes. Many different RdRp enzymes exist, but researchers assumed they were tissue specific and that all cells from the same tissue used the same RdRp enzyme to carry out RNAi.
Jose and his colleagues first genetically modified nematode worms to fluoresce green. Next, they genetically modified the worms to make double-stranded RNA in neurons with the sequence matching the gene for green fluorescence. The double-stranded RNA moved throughout the worms’ bodies and turned off the gene for green fluorescence in many cell types, including intestinal cells.
The researchers then removed the RdRp enzyme thought to be responsible for a critical step in RNAi gene silencing in intestinal cells. Only some of the cells fluoresced green, meaning that something continued to silence the fluorescent gene in some of the other intestinal cells. The RdRp they removed is called RRF-1, but there are three other RdRp’s in the species of worm Jose and his colleagues studied (C. elegans).
Next, the scientists turned to the other three RdRp enzymes present in the nematode and eliminated one at a time to determine which turned off the green fluorescent gene in the worms’ intestines. The culprit: EGO-1, which was thought to aid in RNAi only in reproduction-related germline cells.
Repeating their experiments in multiple individuals revealed that some cells required the RdRp called RRF-1 for silencing, but others did not because they could use EGO-1. Additionally, which intestinal cells used EGO-1 appeared to be random.
“We found that which enzyme gets used varies from cell to cell, and which cells use which enzyme varies from worm to worm,” Jose said. “You can’t tell from looking at them which cells use the alternative molecule, and from the cell’s perspective, it doesn’t matter which molecule it uses.”
According to Jose, functional mosaicism could explain the persistence of some bacterial cells and cancer cells after treatment. If some cells persist by using a different mechanism than the one targeted by therapy, those cells may be the underlying cause of some instances of cancer relapse or antibiotic resistance.
“It may not be that these diseases are developing some new mechanism to survive,” Jose said. “It may be that they already had more than one way of doing the same thing, and the alternates step in to replace whatever the therapeutic treatments knock out.”
Current research into cellular processes is often conducted at the level of body tissues. Scientists combine multiple cells from a given tissue and examine them as a whole with the assumption that cellular processes are the same for all of the cells. According to Jose, therapeutic approaches should now consider the possibility of functional mosaicism, which requires closely examining what’s going on at the single-cell level both before and after treatment.
Jose and his team suggest that functional mosaicism may also provide insight into the evolution of similar traits that develop using different mechanisms.
“Imagine starting off with two equally good ways of doing one thing, and as populations separate and begin to speciate, a mutation could cause one population to get stuck doing that thing one way, and the other population could get stuck doing it the alternative way,” Jose said. “You could end up with two completely different ways of doing the same thing.”
For example, although vision exists throughout the animal kingdom, eyes develop differently between animal groups. Functional mosaicism in a biological process responsible for sensing light in an early ancestor could have led to the divergence in visual systems that exists today among modern animals.
“Now that we know about functional mosaicism, we can go looking for it,” Jose said. “It doesn’t exist in every process to be sure, but it is something we have to be aware of.”
###
Additional co-authors of the research paper from UMD’s Department of Cell Biology and Molecular Genetics include postdoctoral associates Snusha Ravikumar and Sindhuja Devanapally.
This work was supported by the National Institutes of Health (Award No. R01GM111457). The content of this article does not necessarily reflect the views of this organization.
The research paper, “Gene silencing by double-stranded RNA from C. elegans neurons reveals functional mosaicism of RNA interference,” Snusha Ravikumar, Sindhuja Devanapally and Antony M. Jose, was published in the September, 10, 2019, issue of Nucleic Acids Research.
Media Relations Contact: Kimbra Cutlip, 301-405-9463, [email protected]
University of Maryland
College of Computer, Mathematical, and Natural Sciences
2300 Symons Hall
College Park, Md. 20742
http://www.
@UMDscience
About the College of Computer, Mathematical, and Natural Sciences
The College of Computer, Mathematical, and Natural Sciences at the University of Maryland educates more than 9,000 future scientific leaders in its undergraduate and graduate programs each year. The college’s 10 departments and more than a dozen interdisciplinary research centers foster scientific discovery with annual sponsored research funding exceeding $175 million.
Media Contact
Kimbra Cutlip
[email protected]
Original Source
https:/