Researchers develop tunable system that harnesses the spread of cargo carried by gene drives
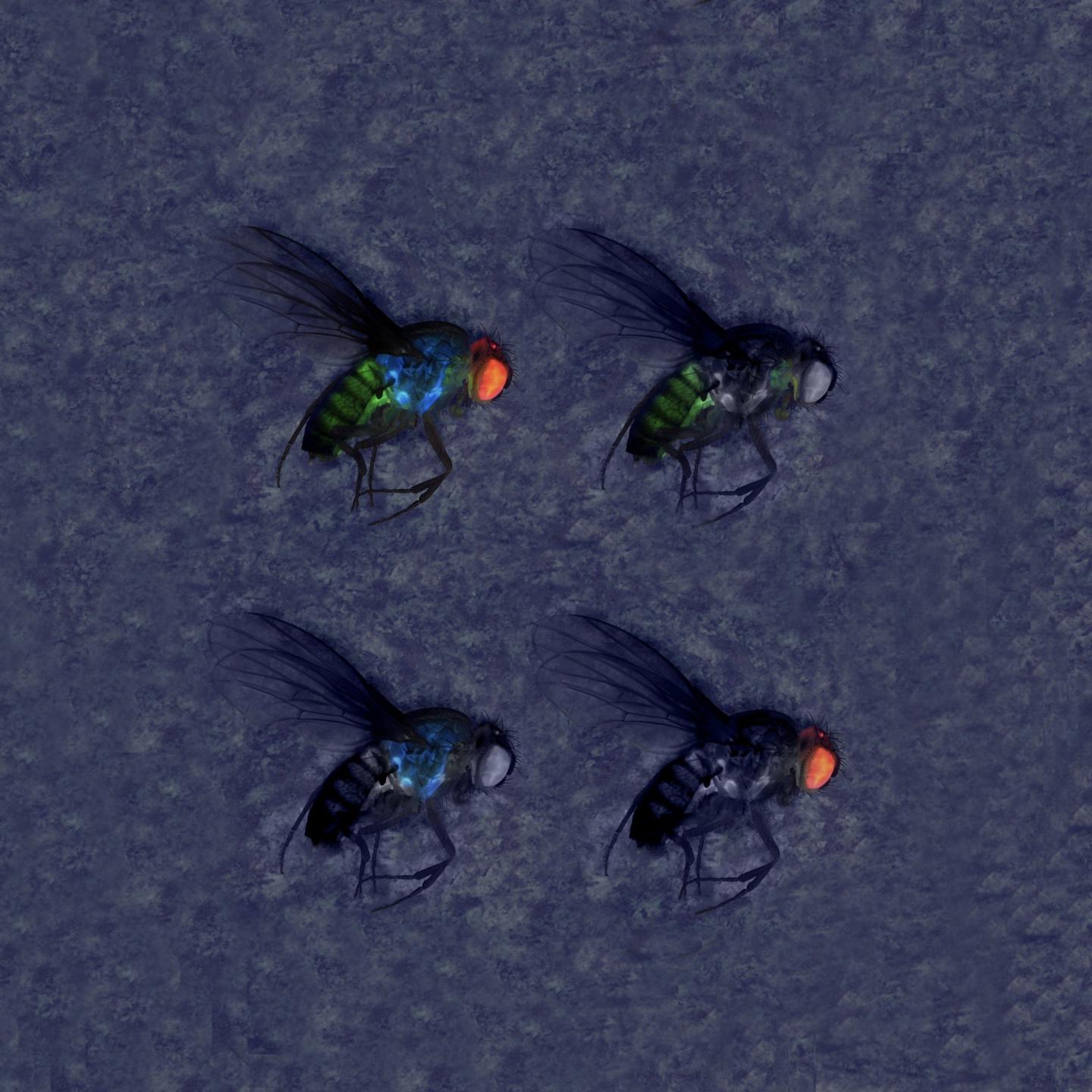

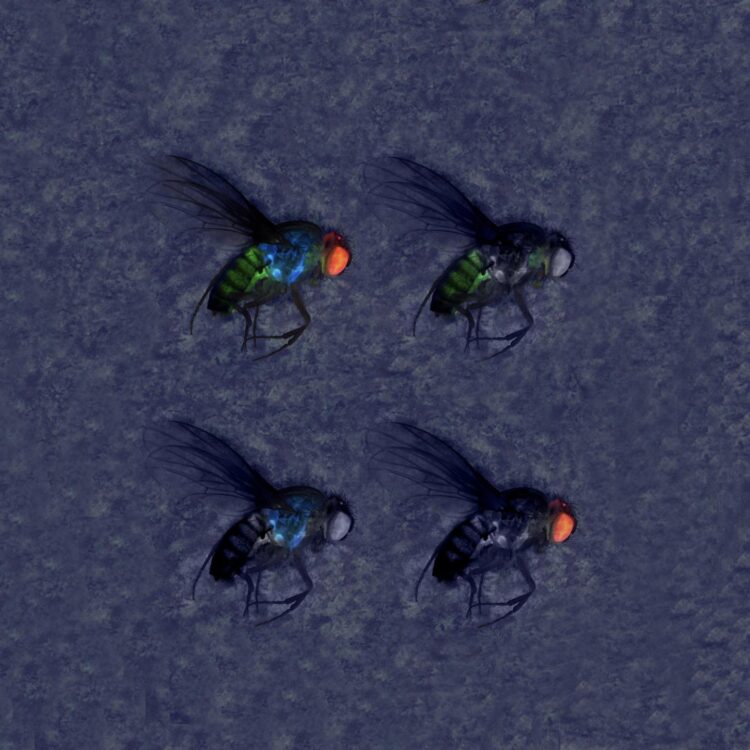
Credit: Gerard Terradas, UC San Diego
Powerful new genetic engineering methods have given scientists the potential to revolutionize several sectors of global urgency.
So-called gene drives, which leverage CRISPR technology to influence genetic inheritance, carry the promise of rapidly spreading specific genetic traits throughout populations of a given species. Gene-drive technologies applied in insects, for example, are being designed to halt the spread of devastating diseases such as malaria and dengue by preventing mosquito hosts from becoming infected. In agricultural fields, gene-drives are being developed to help control or eliminate economically damaging crop pests.
But along with the capacity to alter populations, concerns have been raised regarding the long-term effects of these transformative new technologies in the wild. Researchers and ethicists have voiced questions about how gene drives, once turned loose in a regional population, could be held in check if necessary.
Now, researchers at the University of California San Diego, Tata Institute for Genetics and Society (TIGS) at UC San Diego and their colleagues at UC Berkeley have developed a new method that provides more control over gene drive releases. Details of the new “split drive” are published March 5 in the journals Nature Communications and eLife.
The most common gene drives employ a two-component system that features a DNA-cutting enzyme (called Cas9) and a guide RNA (or gRNA) that targets cuts at specific sites in the genome. Following the Cas9/gRNA cut, the gene drive, along with the cargo it carries, is copied into the break site through a DNA repair process.
While classic gene drives are designed to spread autonomously, the newly developed system is designed with controls that separate the genetic implementation processes. The split-drive system consists of a non-spreadable Cas9 component inserted into one location in the genome and a second genetic element that can copy itself–along with a beneficial trait–at a separate site. When both elements are present together in an individual, an “active gene drive” is created that spreads the element carrying the beneficial trait to most of its progeny. Yet, when uncoupled, the element carrying the beneficial trait is inherited under typical generational genetics rules, or Mendelian frequencies, rather than spreading unrestrained.
As described in the Nature Communications paper, by creating slight fitness costs that eventually eliminate the Cas9 enzyme from the population, the split-drive system vastly increases control and safety of the genetic deployments.
“Studying drives in essential genes is not a novel idea, per se, but we observed that certain split situations were able to spread a cargo effectively upon a first introduction while leaving no trace of Cas9 after a few generations, as well as few mistakes in the DNA repair process that got rapidly diluted out,” said Gerard Terradas, first author in the Nature Communications paper and a postdoctoral scholar in the UC San Diego Division of Biological Sciences.
The Nature Communications paper also spells out advantages on how gene drives are perceived by the public, as efforts to alter wild populations could be flexibly designed in a variety of ways per the desired outcome.
The new split-drive system follows research announced in September in which UC San Diego researchers led the development of two new active genetics neutralizing strategies that are designed to halt or inactivate gene drives released in the wild.
“We hope that the flexible design features we have developed will be broadly applicable by enabling tailored approaches to controlling insect vectors and pests in diverse contexts,” said UC San Diego Distinguished Professor Ethan Bier, senior author of the Nature Communications study and science director for TIGS-UC San Diego.
“These seminal papers reflect a tremendous effort, and fruitful cross-UC collaborations, to demonstrate novel gene drive architectures for mitigating the formation of resistant alleles while providing a safe confinable means for modification of wild populations,” said UC San Diego Associate Professor Omar Akbari, senior author of the eLife study.
###
Authors of the new papers include: Gerard Terradas, Anna Buchman, Jared Bennett, Isaiah Shriner, John Marshall, Omar Akbari and Ethan Bier for the Nature Communications paper; and Nikolay Kandul, Junru Liu, Jared Bennett, John Marshall, and Omar Akbari for the eLife paper.
The research was supported by a DARPA Safe Genes Program Grant (HR0011- 17-2-0047) National Institutes of Health grants (R21RAI149161A, R01AI151004, DP2AI152071, R01GM117321), a Paul G. Allen Frontiers Group Distinguished Investigator Award and a gift from the Tata Trusts in India to TIGS-UC San Diego.
Note: Bier has equity interest in two companies he co-founded: Synbal Inc. and Agragene, Inc., which may potentially benefit from the research results. He also serves on the Synbal’s board of directors and the scientific advisory board for both companies. Akbari is a co-founder, consultant, scientific advisory board member and receives income from Agragene.
Media Contact
Mario Aguilera
[email protected]
Related Journal Article
http://dx.