In a groundbreaking study published in Experimental & Molecular Medicine, scientists have unraveled the complex genetic landscape of hereditary breast cancer, identifying four distinct subtypes defined by varying degrees of genomic instability. This discovery not only deepens our understanding of breast cancer heterogeneity but also opens avenues for precision medicine tailored to the intricate molecular profiles of these malignancies. The research, led by Kim et al., represents a significant leap towards more accurately predicting disease progression and therapeutic responses in patients burdened by inherited breast cancer syndromes.
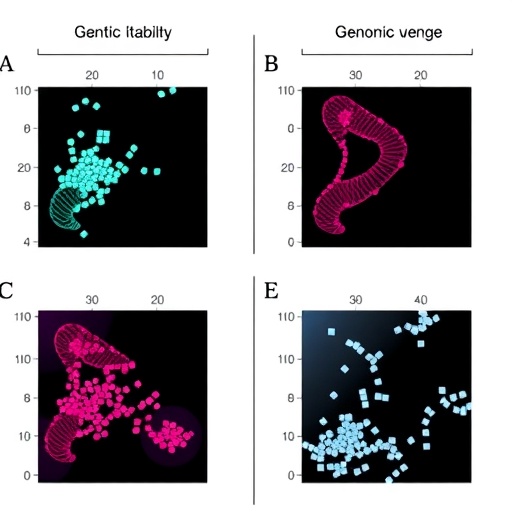
Genomic instability, characterized by the accumulation of mutations and chromosomal aberrations, is a hallmark of many cancers and is particularly prevalent in hereditary breast cancers. However, classifying these tumors based solely on genomic instability levels has proven challenging due to their inherent heterogeneity. Kim and colleagues employed advanced genomic profiling techniques to dissect this complexity, revealing that hereditary breast cancers do not constitute a monolithic group but instead segregate into four subtypes marked by distinct genomic instability patterns and underlying molecular mechanisms.
The study leveraged next-generation sequencing and sophisticated bioinformatic analyses to catalog the genomic alterations across a large cohort of hereditary breast cancer samples. Through comprehensive mapping of single nucleotide variants, copy number changes, and structural rearrangements, the team could stratify tumors according to specific instability signatures. Importantly, these signatures correlated with clinical parameters, suggesting that the identified subtypes bear prognostic and potentially predictive significance.
One of the four subtypes uncovered exhibits relatively low genomic instability but harbors key driver mutations in DNA repair genes. Despite a seemingly stable genome, this subtype presents unique vulnerabilities that could be exploited using targeted therapies aimed at DNA repair pathways. This finding challenges the traditional dogma that high genomic instability is always a prerequisite for aggressive tumor behavior, highlighting the nuanced biology operative even within stable genomes.
Conversely, another subtype demonstrates extensive chromosomal instability characterized by widespread copy number alterations and complex rearrangements. This subtype is associated with aggressive clinical features and poorer outcomes, aligning with current understanding that high genomic chaos often portends treatment resistance and rapid disease progression. Identifying patients belonging to this group could prompt early intervention with novel agents capable of mitigating genome instability-related oncogenesis.
Between these two extremes, the remaining subtypes show intermediate levels of genomic instability, distinguished by specific mutational profiles and epigenetic modifications. The researchers found that each subtype engages distinct cellular pathways to suppress or tolerate genomic damage, underscoring the adaptive plasticity tumors utilize to thrive despite genetic turmoil. These insights lay the foundation for developing subtype-specific therapeutic strategies aimed at disrupting these compensatory mechanisms.
Moreover, the study highlights the importance of integrating genomic instability metrics with other molecular data types such as transcriptomic and epigenomic profiles. This integrative approach enhances subtype discrimination and provides a multidimensional view of tumor biology that transcends single-parameter classification. Such comprehensive profiling could soon become the standard in clinical oncology, facilitating personalized treatment regimens.
Intriguingly, Kim et al. also noted that hereditary breast cancers in carriers of different germline mutations (e.g., BRCA1, BRCA2, PALB2) cluster into distinct genomic instability subtypes. This observation suggests that the inherited mutational background influences tumor evolution and the nature of genomic instability manifesting in the cancer cells. Consequently, genetic counseling and testing may gain additional nuance through consideration of tumor subtype alongside germline variant status.
The implications of subclassifying hereditary breast cancers extend beyond prognostication. For instance, the identification of a subtype with particular susceptibility to PARP inhibitors or immune checkpoint blockade could revolutionize therapeutic paradigms. By aligning treatment modalities with the molecular vulnerabilities delineated in each subtype, clinicians can improve response rates and minimize exposure to ineffective treatments, enhancing patient quality of life.
Further research prompted by this study is likely to focus on validating these subtypes across larger and more diverse populations to ensure generalizability. Additionally, preclinical models tailored to each subtype could accelerate drug discovery efforts and elucidate mechanisms of resistance that arise during treatment. Ultimately, these endeavors will bring the goal of truly personalized medicine within reach for hereditary breast cancer patients.
Another facet of the work includes potential biomarker development based on genomic instability signatures. Non-invasive assays detecting circulating tumor DNA or other components reflective of subtype-specific instability could assist in early diagnosis, monitoring treatment response, and detecting minimal residual disease. This may prove particularly valuable in hereditary cancer syndromes where lifelong surveillance is required.
The study’s methodological advancements also merit attention. The combined application of multi-omics data integration, machine learning algorithms for subtype prediction, and rigorous statistical validation sets a high bar for future cancer genomics research. This integrative framework is poised to be adapted for studying genomic instability in other hereditary and sporadic cancers, fostering a new era of comprehensive precision oncology.
Importantly, this research sheds light on the evolutionary dynamics of breast tumors developing in the context of inherited genetic predisposition. It illustrates how selective pressures and DNA damage repair deficiencies converge to sculpt distinct genomic instability landscapes that ultimately dictate tumor behavior. Understanding these dynamics is essential for crafting interventions that outpace cancer’s ability to adapt and resist therapy.
As knowledge about genomic instability deepens, collaborations between molecular biologists, clinicians, and computational scientists will become ever more crucial. This multidisciplinary synergy will accelerate the translation of findings like those of Kim et al. into tangible improvements in patient care, bringing personalized oncology from bench to bedside with unprecedented precision and efficacy.
In conclusion, the delineation of four genomic instability-based subtypes in hereditary breast cancers marks a paradigm shift in the characterization and management of these diseases. By elucidating the heterogeneity that underpins tumor development and progression, this landmark study empowers clinicians with new tools for tailoring therapies, refining prognoses, and ultimately improving outcomes for women battling hereditary breast cancer worldwide.
Subject of Research: Genomic instability and heterogeneity in hereditary breast cancer subtypes
Article Title: Delineation of the heterogeneity underlying genomic instability in hereditary breast cancers reveals four disease subtypes
Article References:
Kim, S., Lee, S., Kim, H. et al. Delineation of the heterogeneity underlying genomic instability in hereditary breast cancers reveals four disease subtypes. Experimental & Molecular Medicine (2026). https://doi.org/10.1038/s12276-026-01693-4
Image Credits: AI Generated
DOI: 16 April 2026
Tags: bioinformatics in cancer researchbreast cancer heterogeneitycancer genomic alterations analysischromosomal aberrations in cancergenetic mutations in breast cancergenomic instability in breast cancerhereditary breast cancer subtypesinherited breast cancer syndromesmolecular profiling of breast cancernext-generation sequencing breast cancerprecision medicine in oncologytherapeutic targets in hereditary breast cancer