Partial unfolding of proteins may affect their stability and is a challenge in the industry. So how does a cavity destabilize a protein? Would such a cavity be empty? These are questions that researchers from Aarhus University answer in a new study
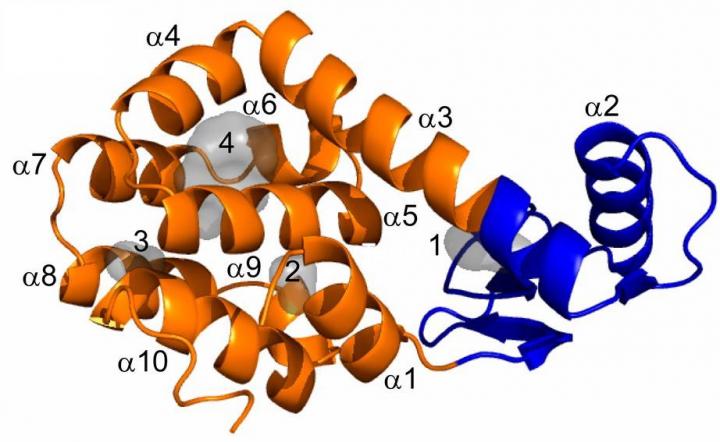
Credit: Proc Natl Acad Sci U S A, copyright 2019 National Academy of Sciences.
Proteins exist as groups of microscopic configurations, regulated by a landscape of free energy, in which there is a multitude of “excited” states that co-exists with the minimum energy structure. These alternatively folded and partially “disordered” states occur continuously due to protein dynamics and are key elements required to understand the function and stability of proteins.
Because these excited states exist only briefly and are lowly populated they are “invisible” to most experimental methods. However, recent developments in NMR spectroscopy allow for their detection and structural investigation at atomic resolution.
In this study, the researchers used a classic model system for protein folding, the L99A mutant of T4 lysozyme. This protein has a cavity in its hydrophobic core that is large enough to fit a benzene ring (a chemical compound consisting of a ring of 6 carbon atoms).
READ ALSO: New efficient method for urine analysis may tell us more
Pressure reveals invisible states
Mulder and his team used Nuclear Magnetic Resonance (NMR) spectroscopy coupled with hydrostatic pressure to monitor “invisible” excited states. High pressure favors compact states, and the protein unfolds or collapses at high pressure to remove cavities.
The researchers have succeeded in obtaining a unique picture of the hierarchy of unfolded states in the protein’s energy landscape by subjecting it to pressure. Furthermore, with these pressure perturbations, they have been able to identify empty protein cavities and determine the energetic consequences of filling these with water.
Partial unfolding of unstable parts of proteins is a major concern in the development of industrial enzymes and biological drugs, as well as a starting point for protein deposition diseases. The approach shown in this study here establishes a powerful way to rationally understand and gain control of protein stability at the atomic level.
READ ALSO: How good are protein disorder prediction programmes actually?
###
More information
The work is financially supported by the Japan Society for the Promotion of Science, Danish Ministry of Higher Education and Science and the Carlsberg Foundation.
The research was carried out by researchers from Interdisciplinary Nanoscience Center (iNANO) and Department of Chemistry, and Department of Molecular Biology and Genetics
at Aarhus University (AU), in collaboration with researchers from Graduate School of Life Sciences and College of Pharmaceutical Sciences (Japan). Associate Professor Frans Mulder (iNANO and Department of Chemistry, AU) was in charge of the research team.
Read more about the results in Proceedings of the National Academy of Sciences:
Xue, M.; Wakamoto, T.; Kejlberg, C.; Yoshimura, Y.; Nielsen, T. A.; Risør, M. W.; Sanggaard, K. W.; Kitahara, R.; Mulder, F. A. A. How internal cavities destabilize a protein. Proc Natl Acad Sci U S A. 2019. DOI: 10.1073/pnas.1911181116.
Contact
Associate Professor Frans Mulder
Email: [email protected]
Tel: +4587155889
Media Contact
Frans Mulder
[email protected]
Original Source
http://inano.
Related Journal Article
http://dx.




