New CRISPR-Cas9 variants respond to viral and cellular proteases
Credit: UC Berkeley
CRISPR-Cas9 is a revolutionary tool in part because of its versatility: created by bacteria to chew up viruses, it works equally well in human cells to do all sorts of genetic tricks, including cutting and pasting DNA, making pinpoint mutations and activating or inactivating a gene.
University of California, Berkeley, researchers have now made it even more versatile by giving it an “on” switch, allowing users to keep the Cas9 gene editor turned off in all cells except its designated target.
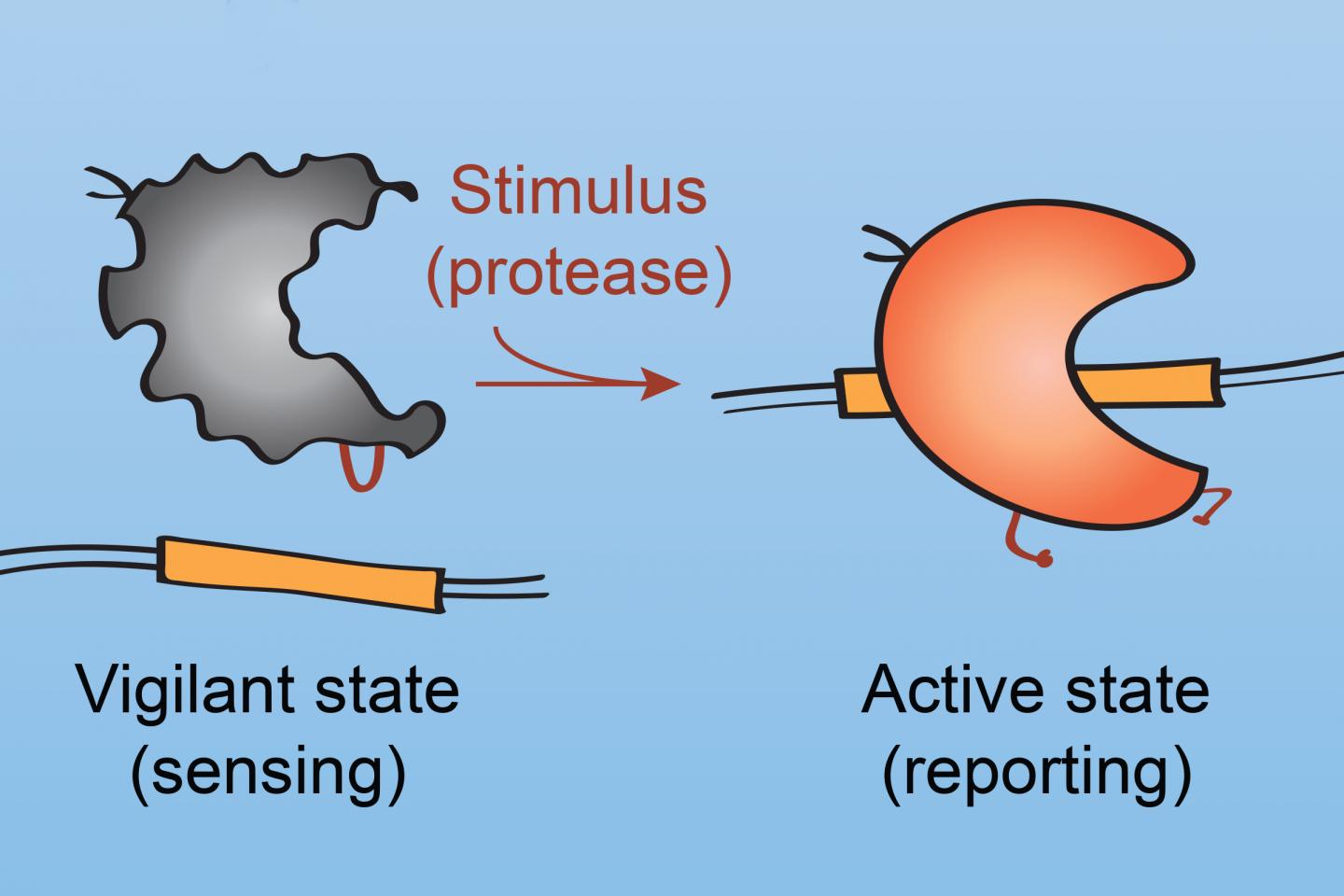
The redesigned Cas9 enzyme — which the researchers refer to as ProCas9 — is fully functional except for containing a length of protein that needs to be snipped before the enzyme can bind and cut DNA. If scientists insert a short length of protein that can be cut only by a particular enzyme, such as one used exclusively by cancer cells or an infectious virus or bacteria, that enzyme becomes a trigger to turn on Cas9.
ProCas9 essentially “senses” the type of cell it’s in based on the protein-cutting enzyme — a so-called protease — present.
“This is an extra layer of security you could put on the molecule to ensure accurate cutting,” said David Savage, a UC Berkeley associate professor of molecular and cell biology.
It also endows the Cas9 protein with programmable inputs in addition to its programmable outputs.
“There are a lot of proteases that regulate signaling pathways in cells, transform normal cells into cancer cells, and are involved in pathogen infection,” said Savage. “If we can sense these signals, we can tap into and respond accordingly to these important pathways.”
In the study, Savage and his colleagues demonstrated protease control of Cas9 by making it sensitive to both plant and human viruses, such as West Nile Virus. In the future, they believe this sort of technology could be used to import the CRISPR-Cas9 bacterial immune system into plants to help them fend off damaging viral pathogens.
The study by researchers from UC Berkeley and the Gladstone Institutes in San Francisco will be published online Jan. 10 in the journal Cell.
Stripping Cas9 to its essentials
Savage’s original research goal was to pare down the Cas9 protein — the “scissors” that actually snip DNA to do gene editing — to its simplest parts, to get the most robust gene editor possible. The simpler it is, the easier it is to work with and to ferry into cells.
“We know the Cas proteins are complex and that they have all sorts of regulation that is critical to how they function in a bacterial immune system,” he said. “The broad goal of our work is to tame them for use in humans and to strip away the unnecessary things not relevant for genome editing.”
In particular, he wanted to redesign Cas9 so it would be easier to attach other proteins. This would allow Cas9 to carry proteins with a variety of functions to the right spot on DNA. These are known as fusion proteins, promising variants of which can alter gene expression or, in the case of a technology known as base-editing, alter one base or nucleotide in the DNA with pinpoint accuracy.
The technique he used to reengineer Cas9, called circular permutation, has never been tried on a protein as complex as Cas9. Circular permutation involves taking the amino-acid string of the Cas9 protein and cutting it, switching the order of the two segments, and then allowing it to fold into a new 3D configuration. He did this for all possible ways of cutting the protein into two pieces.
While you might think this would completely wreck the protein, in about 10 percent of cases, the new protein still worked, as if he had merely rearranged the protein’s subunits in a different way that did not affect how they functioned. This may work because, as CRISPR-Cas9 inventor Jennifer Doudna and her colleagues have shown, the Cas9 protein complex is highly flexible and moves around as it grabs onto guide RNA, binds to DNA and moves into position to cut DNA strands.
Savage is currently exploring some Cas9 rearrangements that may provide a better scaffold for fusion proteins, bringing them closer to the DNA strand they are targeting.
In the process of rearranging the Cas9 scaffold, he found, serendipitously, that the way in which he reconnected the two protein segments made a difference.
“When we cut the protein and moved the old piece to a new place within the protein, the system became very sensitive to how you linked the two fragments together,” he said. “We realized that we could use that sensitivity to engineer the protein to have protease recognition sites.”
The study shows that “we’re not stuck with what nature gave us with regards to genome-editing proteins,” he said. “These proteins can be elaborately optimized and turned into scaffolds not found in nature but possessing the right properties for use in human cells.”
###
Savage’s UC Berkeley co-authors include Doudna, a professor of molecular and cell biology and a Howard Hughes Medical Institute investigator, and Gladstone Institutes postdoctoral colleague Christof Fellmann; Adam Arkin, a professor of bioengineering; and Benjamin Oakes, Harneet Rishi, Kian Taylor, Shawn Ren, Dana Nadler and Rayka Yokoo of UC Berkeley and the Innovative Genomics Institute. Co-first authors Oakes and Fellmann are IGI Entrepreneurial Fellows. Most of the work was supported by the National Institutes of Health (1DP2EB018658-01).
Media Contact
Robert Sanders
[email protected]
510-643-6998