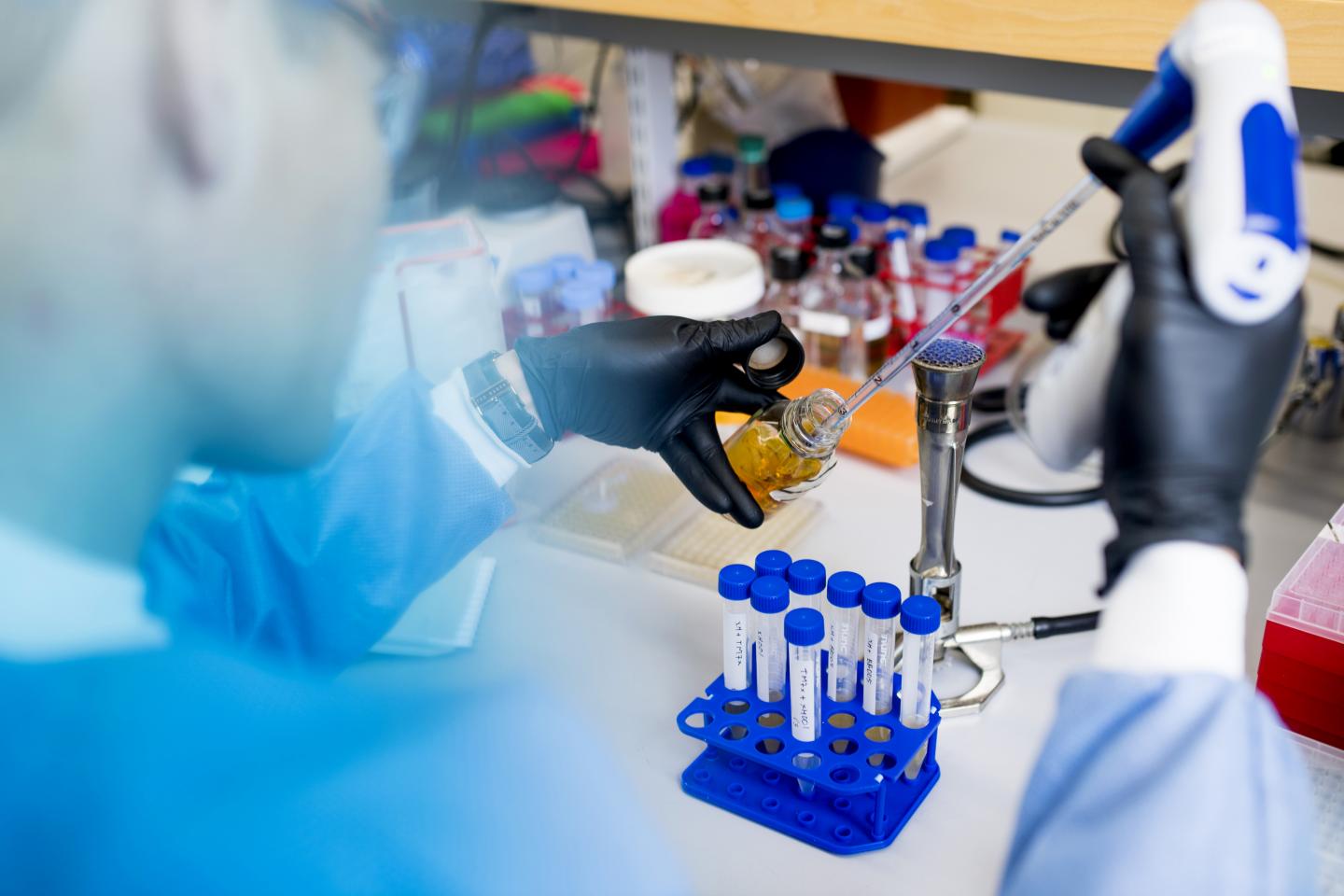
Credit: Photo by Matthew Modoono for Forsyth Institute
Inside your mouth right now, there is a group of bacteria whose closest relatives can also be found in the belly of a moose, in dogs, cats, and dolphins, and in groundwater deep under the Earth’s surface. In a stunning discovery, scientists have found that these organisms have adapted to these incredibly diverse environments–without radically changing their genomes.
The organisms are members of the TM7, or Sacchraribacteria, phylum. These are ultra-small, parasitic bacteria with small genomes that belong to a larger group called the Candidate Phyla Radiation (CPR). These CPR bacteria are mysterious “dark matter” that represent more than 25 percent of all bacterial diversity, yet we know very little about them since the vast majority remain uncultivated.
In research first published as a pre-print in 2018, and now formally in the journal Cell Reports, scientists describe their findings that Saccharibacteria within a mammalian host are more diverse than ever anticipated. The researchers also discovered that certain members of the bacteria are found in the oral cavity of humans, the guts of other mammals, and in groundwater. While these environments are all very different, the bacteria’s tiny genomes remain minimally changed between humans and groundwater. This indicates that humans acquired the bacteria more recently, on an evolutionary timescale.
“It’s the only bacteria we know that has hardly changed when they adapted to humans,” said Dr. Jeffrey Scott McLean, a microbiologist and Associate Professor of Periodontics at the University of Washington School of Dentistry, and lead author of the paper.
The TM7 bacteria were a complete mystery to scientists until Dr. Xuesong He, Associate Member of Staff at the Forsyth Institute and co-author of the paper, first isolated the bacterium TM7x, a member of CPR, in 2014. Since then, researchers have learned that CPR includes a huge number of different bacteria, all with tiny genomes. These bacteria need a host to survive and are unique in that they can’t make their own amino acids and nucleotides, which are essential building blocks for life.
“I see this as a huge discovery,” said Wenyuan Shi, CEO and Chief Scientific Officer at the Forsyth Institute and co-author on the paper. “This creature survives in both humans and groundwater, which indicates there are similarities that allow these bacteria to adapt to humans.”
Previous research by Dr. Batbileg Bor, Assistant Member of Staff at the Forsyth Institute and co-author of the paper, showed that TM7 can easily jump from one bacterial host to another. This could explain how they ended up in mammals, since mammals drink groundwater.
“The most likely reason we see a large diversity of these bacteria in humans, yet one group of bacteria remains nearly identical to those in groundwater, is that some groups were acquired in ancient mammal relatives and they expanded over time across mammals, whereas this one highly similar group more recently jumped directly into humans,” McLean said.
TM7 and other ultra-small, parasitic bacteria within CPR may play important roles in health and disease that we have yet to discover. Since they act as parasites–living with and killing other bacteria–TM7 could change the overall microbiome by modulating the abundance of bacteria, McLean said. Scientists are just scratching the surface of understanding how much our microbiome impacts our overall health.
Another major contribution of this research has been developing a systematic way to name these newly discovered bacteria, setting the foundation for classifying other isolated strains in the future.
The fact that humans acquired TM7 recently is a discovery that has broader implications for understanding our co-evolutionary pathways with the microbes that live on and within us.
“There are only a couple hundred genes that are different in these ultra-small bacteria between what lives deep in the subsurface environment and those that have become common bacteria in our mouths,” McLean said. “That is a remarkable feat for bacteria missing so many genes.”
###
Media Contact
Alexandra Nicodemo
[email protected]
Original Source
https:/
Related Journal Article
http://dx.