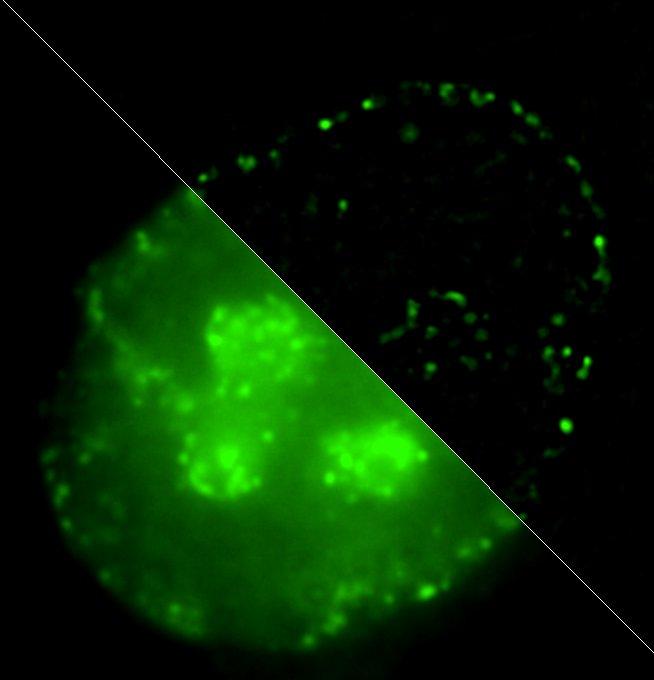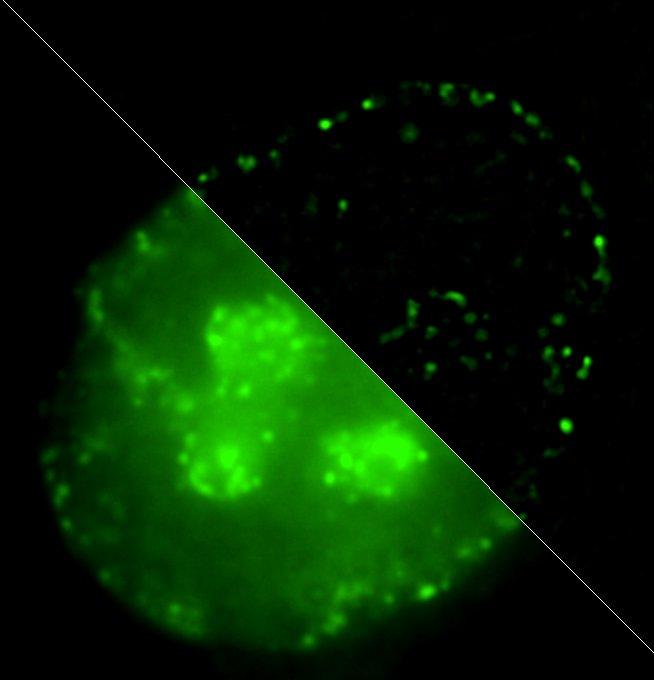
Credit: Igor Kireev
Scientists from the Lomonosov Moscow State University in collaboration with their colleagues from the USA have conducted a research on DNA packaging in the cell nucleus and its alteration in the process of DNA replication. DNA packaging is supposed to have strong influence on genes' activity. It's also one of the mechanisms of epigenetic control of gene expression.
Igor Kireev, Doctor of Biological Sciences, who is the Chief of Department of electron microscopy at the Belozersky Institute of Physico-Chemical Biology and one of the article authors, says: "Although all the cells in a given organism possesse similar genetic information, not all genes operate in a particular cell type. In fact, the set of active genes determines the cell fate. There are some cellular mechanisms assisting the cell to remember, which genes should be activated and which should not. This is the epigenetic control or, in other words, cell memory, that determines what the cell is and due to which genes it has been formed in such a way. We were interested in so called high levels of chromosome structural organization, which are formed through a series of consecutive phases in DNA packaging."
DNA exists in a cell as a complex of proteins — chromatin. Initial stages of chromatin compaction are studied quite well. These are nucleosomes, protein globules, each is around 10 nanometers long and consisting of 8 histone protein molecules, around which DNA is wrapped. Then a chain of nucleosomes is packed in a way that we don't know yet, resulting in thicker fibrils and chromonemata. At the end we get a very high degree of compaction. So, the length of a most densely compacted mitotic chromosome is 20 000 times smaller than the length of DNA, which is packed inside it.
Replication is the process of copying DNA molecule and transcription is the process of RNA synthesis, both using DNA as a template. It was traditionally assumed high compaction of DNA in chromatin to be a hindrance for these template-based synthesis processes and should be disassembled on a scale of quite large chromatin domains. It was fairly difficult to identify these domains and analyze their structural organization at high spatial resolution without disturbing their natural structure.
Igor Kireev comments: "We've offered a method of labeling chromatin domains based on differences in replication timing of active and inactive gene loci. While actively transcribed chromosome regions – so called euchromatin – replicate at the very beginning of synthetic phase of cell cycle, inactive "silent" heterochromatin replicates during its second half. Our technique has allowed to perform nondestructive detection of these two fractions of chromatin at electron microscopy level. It's based on combination of new-synthesized DNA labeling using click-chemistry methods with subsequent detection of reaction products by correlative siperresolution optical microscopy and immunoelectron microscopy. In other words, we can see one and the same molecule in this cell both in optical and electron microscopes."
Application of this approach has given two surprising observations. First of all, "working" chromatin, contrary to traditional ideas, maintains quite high level of packaging, as it's represented by highly structured chromatin fibrils of lhigher order – chromonemata. Secondly, it's turned out that DNA in chromonema structure possesses high degree of structural flexibility, what means that it's able to sort of flow from one section of chromonema fiber to the neighboring one. However, the general dense structure of chromonema doesn't change. These observations don't fit the existing theories of chromosome spatial organization, however, at the same time allow to offer new hypotheses concerning mechanisms of epigenetic information transfer in the process of cell division.
Igor Kireev continues: "We've made an assumption, that newly-synthetized DNA in chromatin structures isn't fixed but could move inside them, interacting with that DNAsegments, which hasn't doubled yet and "remembers everything". Moreover, this DNA contains necessary molecular components for recovery of lost epigenetic information".
Another result, mentioned in the article, which has been published in Current Biology, lies in the fact that genome structural organization doesn't have strict hierarchy. There are, of course, some gradual levels of DNA organization. It was thought earlier that there should be a strictly ordered system, which uniquely leads to chromosome formation. But now it's turned out that everything could be organized in a different way – there are some building principles, but within specified chromatin frameworks DNA has some freedom and flexibility.
Igor Kireev shares future plans: "Further research development includes, first of all, transition to analysis of individual chromosome loci. We are going to label them both on optical and electron microscopic levels with the help of TALE technology. And, secondly, further research will tackle with elaboration of even more native methods of in vivo labeling, which are compatible with such advanced technologies, as cryo-electron microscopy".
Scientists hope to approach closely to DNA spatial organization, applying direct methods of visualization of chromatin packaging ways with high resolution. As far as practical part is concerned, the researches will help to clarify structural aspects of epigenetic control of gene expression and, probably, offer some ways of its regulation. This will contribute to elaboration of more effective approaches to therapeutic intervention, for instance, to fight cancer or ageing, where epigenetic component plays an important role.
###
Media Contact
Vladimir Koryagin
[email protected]
http://www.msu.ru