Credit: LIU Yang
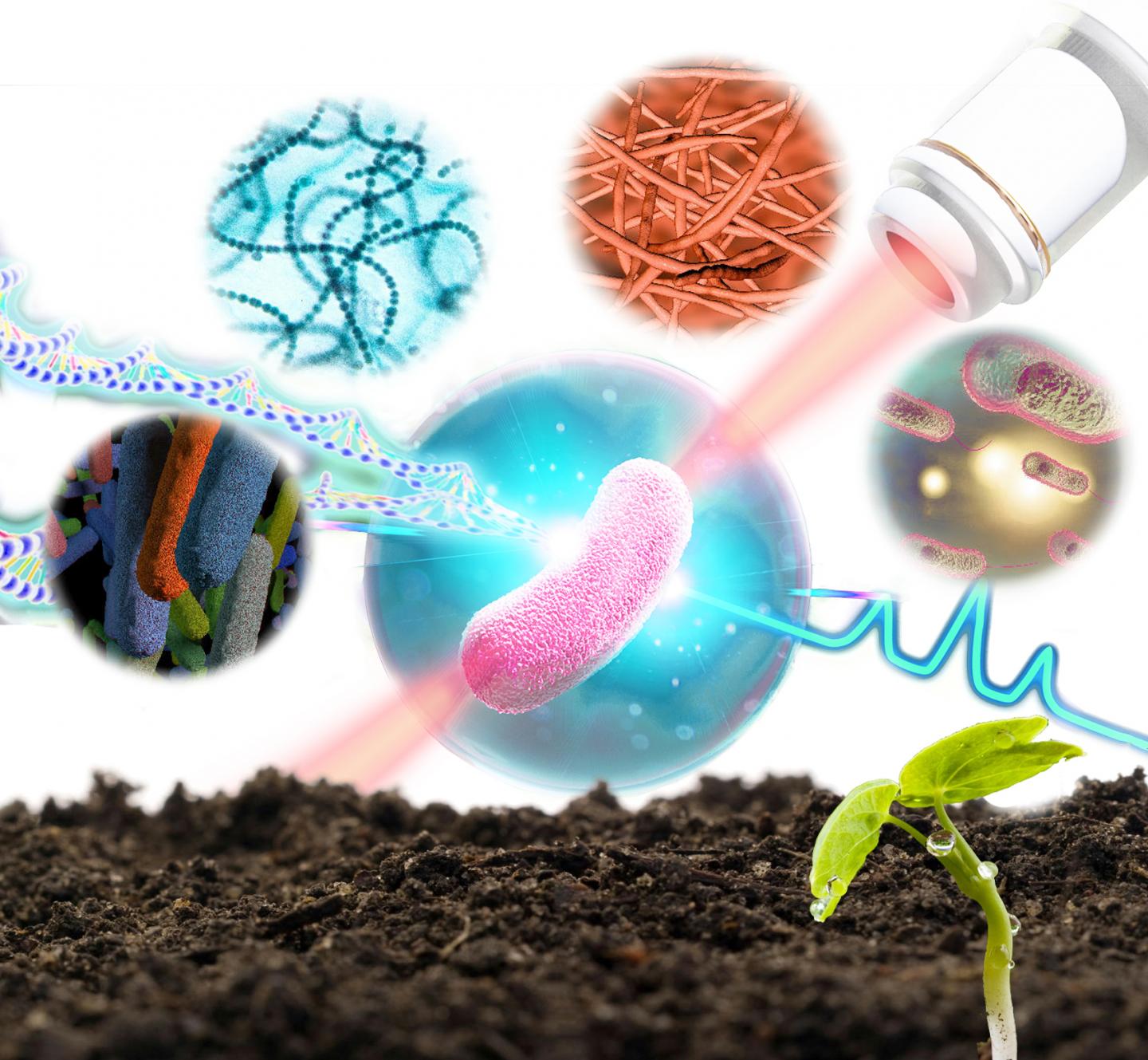
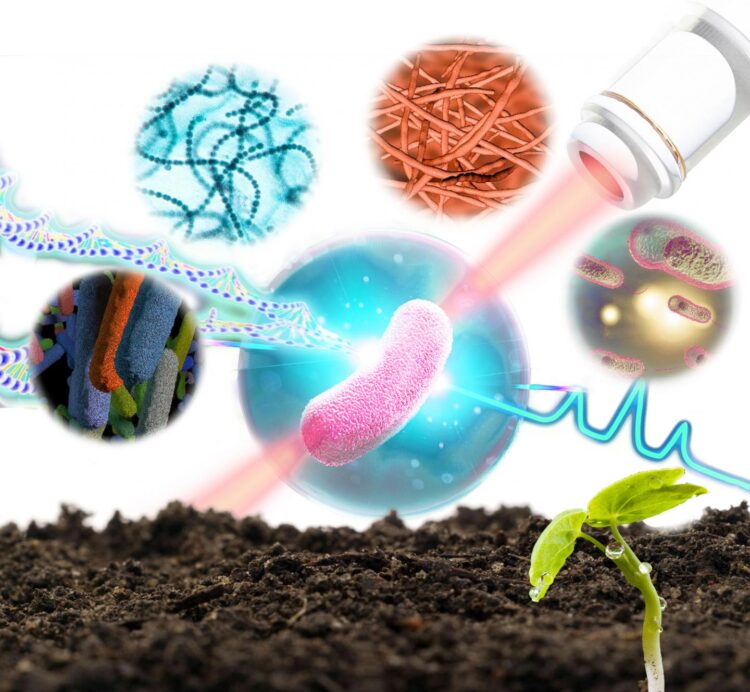
Researchers from the Single-Cell Center at the Qingdao Institute of Bioenergy and Bioprocess Technology (QIBEBT) of the Chinese Academy of Sciences have developed a technique to sort and sequence the genome of bacteria in soil one bacterial cell at a time, while also identifying what its function is in the soil environment.
Their study was published in the journal mSystems on May 27.
Soil is home to a vast and complex microbiome, which features arguably the highest genomic diversity and widest heterogeneity of metabolic activities of cells on Earth. In turn, these metabolic activities can in principle provide the foundation for industrial production of numerous compounds of value.
The ability to pinpoint “who is doing what” in the soil microbiome has until now been extremely difficult at the resolution of a single cell, as normally a great many cells, typically millions of them, have had to be analyzed at the same time.
In 2020, researchers from the QIBEBT Single-Cell Center developed a bacteria-profiling technique called Raman-Activated Gravity-driven single-cell Encapsulation and Sequencing, or RAGE sequencing. This sequencing technique uses laser “tweezers” and takes advantage of the properties of gravity to permit analysis of bacteria cells one by one. The form and structure of a bacterium are then investigated using ‘Raman spectroscopy’, a method of analysis that uses how light interacts with the chemical bonds in a molecule to enable identification.
They have now developed their RAGE sequencing technique into a new scientific instrument they are calling a Raman-activated Cell Sorter-Sequencer or RACS-Seq. It is the first instrument in the world that can pinpoint “who is doing what” at precisely one-cell resolution in complex ecosystem. And they applied it to the bacteria in soil.
The Raman spectra of a single cell offers an intrinsic biochemical fingerprint, and this in turn can be used as a proxy of its metabolic activity. When this single-cell Raman spectra (SCRS) is coupled with probing of the isotopes involved in this activity, the technique can reveal the amount of chemical from the cell’s environment that it is taking up.
Isotopes are often used as a way to track or trace chemical processes. Deuterium, for example, is an isotope of hydrogen with one neutron (regular hydrogen has zero neutrons), and is found in heavy water (water molecules where the regular hydrogen is replaced by deuterium: D2O instead of H2O).
By feeding the bacteria with heavy water, the general metabolic activity of the cells can be tracked via Raman spectra. The technique can also detect not only uptake of D2O but also different isotopes of carbon or nitrogen. This can even reveal the particular cellular profile of synthesis of biological molecules.
This metabolic tracking can then be linked via the RAGE technique to individual cells whose genome can be sequenced.
“This should allow researchers to ‘mine’ soil to find bacteria of interest that are in the business of producing particular carotenoids, lipids, polysaccharides, protein and even antibiotics,” said JING Xiaoyan, a microbiome scientist with the QIBEBT Single-Cell Center and the article’s lead author.
The researchers now want to further improve the throughput of their technique, at a few cells per minute currently, to speed up and fully automate the process.
“Our goal ultimately is to keep optimizing the RACS-Seq instrument, so that it becomes a universal and highly versatile tool for precisely probing target cells or metabolic activities of interest,” added XU Jian, the article’s corresponding author and Director of the QIBEBT Single-Cell Center. “And not just from soil, but from any other complex natural ecosystem.”
###
Media Contact
CHENG Jing
[email protected]
Original Source
https:/