Credit: APSB
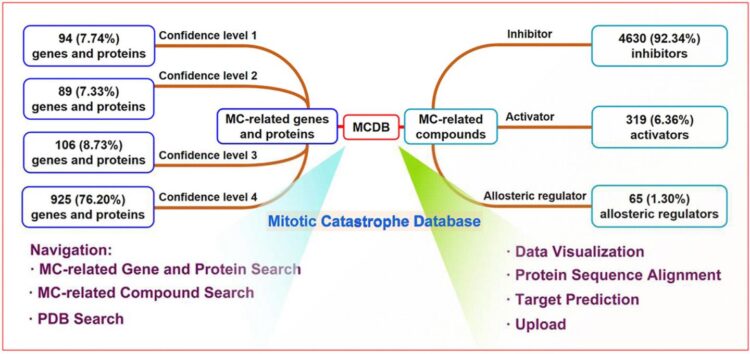
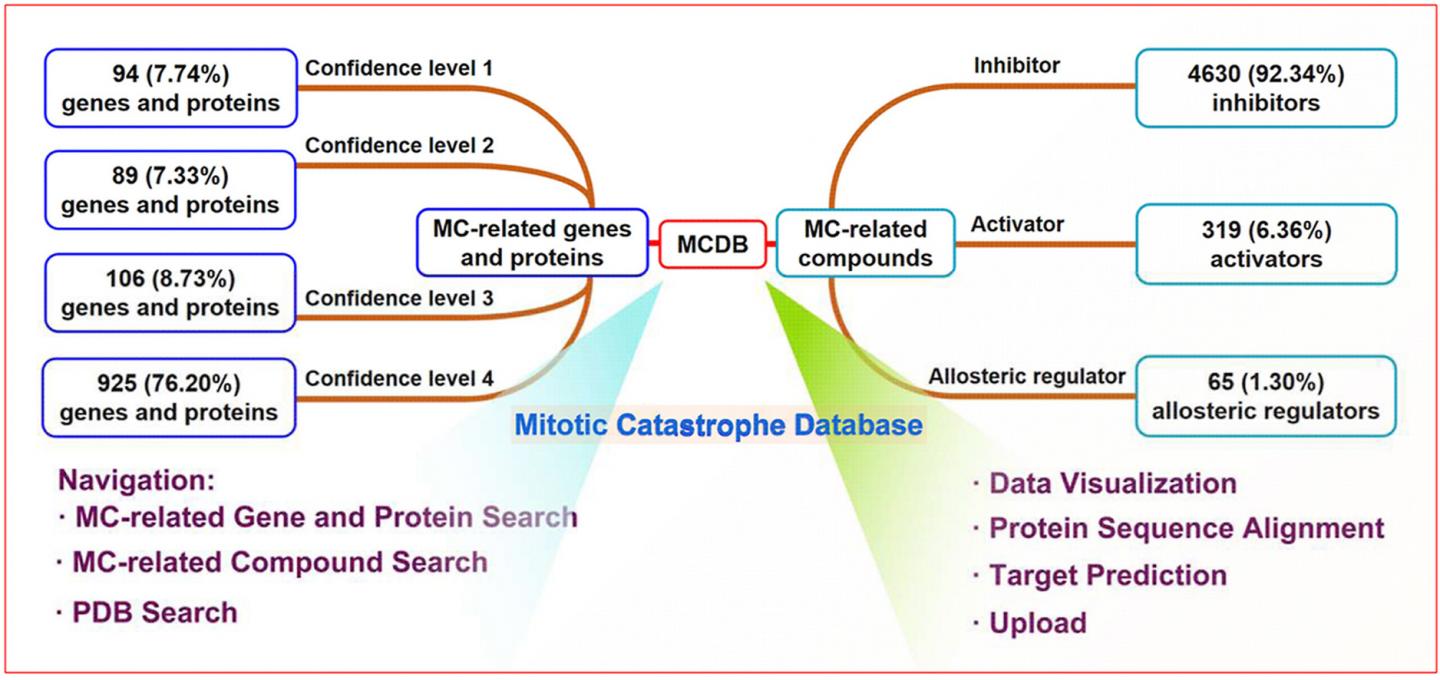
Mitotic Catastrophe Database (MCDB) is a proprietary, standard, and comprehensive database for Mitotic catastrophe (MC) related data facilitating the exploration of MC for all researchers in the fields of medicinal chemistry, molecular biology, bioinformatics and oncology.
MC is a form of programmed cell death induced by mitotic process disorders, which is important in tumor prevention, development, and drug resistance. As availability of MC data underpins tumor-related biomedical and clinical studies, the development of a professional and comprehensive database to curate MC-related data is a matter of increasing priority.
MCDB consists of 1214 genes/proteins and 5014 compounds collected and organized from more than 8000 research articles. MCDB defines the confidence level, classification criteria, and uniform naming rules for MC-related data, which greatly improves data reliability and retrievability. MCDB also develops protein sequence alignment and target prediction functions. The former can be used to predict new potential MC-related genes and proteins, and the latter can facilitate the identification of potential target proteins of unknown MC-related compounds.
###
MCDB is available at http://www.
Article reference: Le Zhang, Lei Zhang, Yue Guo, Ming Xiao, Lu Feng, Chengcan Yang, Guan Wang, Liang Ouyang, MCDB: A comprehensive curated mitotic catastrophe database for retrieval, protein sequence alignment, and target prediction, Acta Pharmaceutica Sinica B, 2021, ISSN 2211-3835, https:/
Keywords: Mitotic catastrophe; Database; Protein sequence analysis; Target prediction; Data mining
The Journal of the Institute of Materia Medica, the Chinese Academy of Medical Sciences and the Chinese Pharmaceutical Association.
For more information please visit https:/
Editorial Board: https:/
APSB is available on ScienceDirect (https:/
Submissions to APSB may be made using Editorial Manager® (https:/
CiteScore: 12.5
Impact Factor: 11.413
ISSN 2211-3835
Media Contact
Morgan lyons
[email protected]
Related Journal Article
http://dx.