Credit: DICP
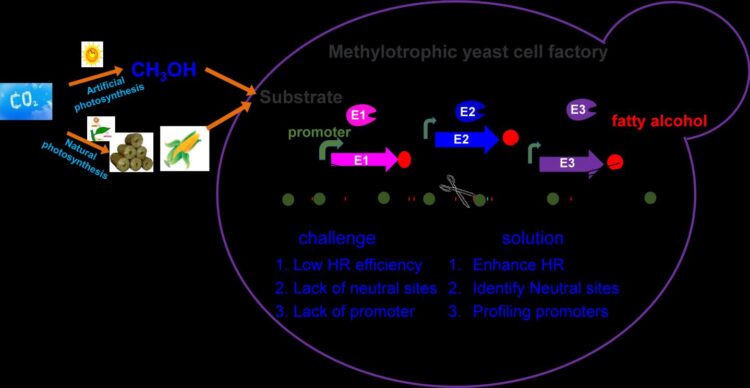
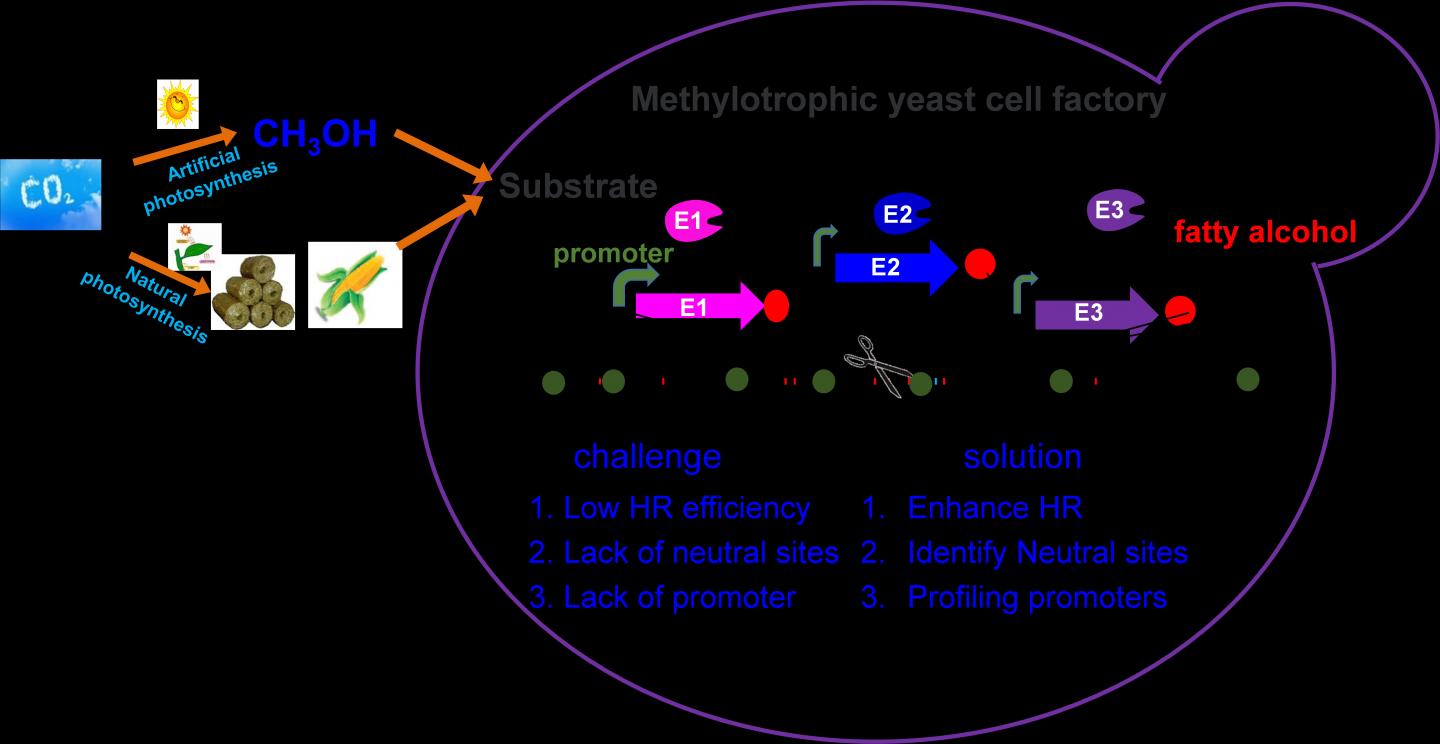
Pichia pastoris (syn. Komagataella phaffii), a model methylotrophic yeast, can easily achieve high density fermentation, and thus is considered as a promising chassis cell for efficient methanol biotransformation. However, inefficient gene editing and lack of synthetic biology tools hinder its metabolic engineering toward industrial application.
Recently, a research group led by Prof. ZHOU Yongjin from the Dalian Institute of Chemical Physics (DICP) of the Chinese Academy of Sciences established an efficient genetic engineering platform in Pichia pastoris.
The study was published in Nucleic Acids Research on July 1.
The researchers developed novel genetic tools for precise genome editing in Pichia pastoris by enhancing homologous recombination (HR) rates and engineering the multiple intrusion-induced rearrangement (MIR) processes. The key gene RAD52, which played crucial role in HR repair in Pichia pastoris, was overexpressed for improving the efficiency of single gene editing to 90%.
Furthermore, they increased the efficiency of multi-fragment recombination at one site by 13.5 times, and identified and characterized 46 neutral sites and 18 promoters for genome integration and gene expression.
Finally, they developed a two-factorial regulation system for regulating fatty alcohol biosynthesis in Pichia pastoris from different carbon sources.
“This advanced gene editing systems can theoretically realize stable loading of more than 100 exogenous genes and precise regulating of gene expression in Pichia pastoris, which will provide convenience for the synthetic biology research of Pichia pastoris. It also provides insights for metabolic engineering of other unconventional yeast,” said Prof. ZHOU.
###
This study was supported by the National Natural Science Foundation of China and Dalian Science and Technology Innovation Funding.
Media Contact
Jean Wang
[email protected]
Original Source
https:/
Related Journal Article
http://dx.