Credit: Bruce Smith
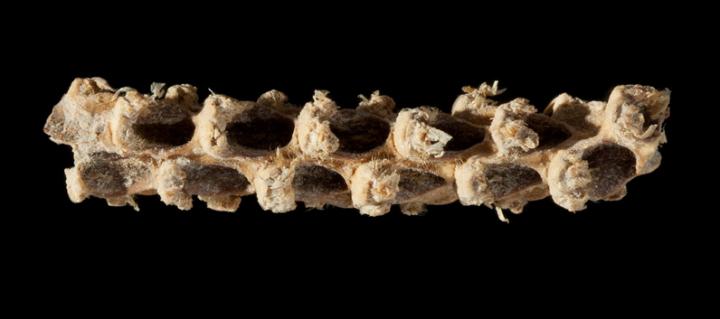
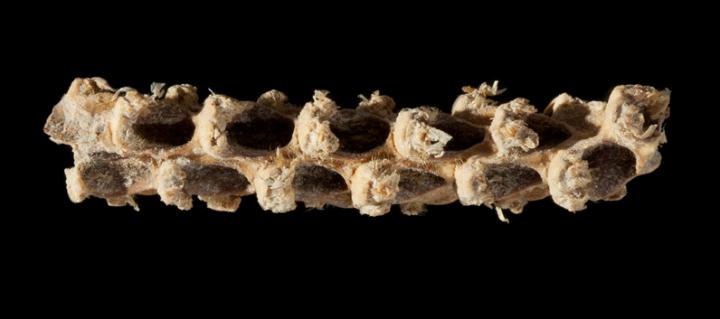
Researchers who have sequenced the genome of a 5,310-year-old corn cob have discovered that the maize grown in central Mexico all those years ago was genetically more similar to modern maize than to its wild ancestor. For example, the ancient maize already carried genetic variants responsible for making kernels soft, a common feature of modern corn. The findings are reported in Current Biology on November 17.
"Around 9,000 years ago in modern-day Mexico, people started collecting and consuming teosinte, a wild grass," says Nathan Wales of the Natural History Museum of Denmark. "Over the course of several thousand years, human-driven selection caused major physical changes, turning the unproductive plant into modern maize, commonly known as corn. Maize as we know it looks so different from its wild ancestor that a couple of decades ago scientists had not reached a consensus regarding the true ancestor of maize."
To better understand the domestication history of the world's most produced crop, Wales and his colleagues, including Jazmín Ramos-Madrigal, sequenced the genome of a 5,310-year-old maize cob from central Mexico. The cob, known as Tehuacan162, was excavated from a cave in the Tehuacan Valley in the 1960s, during a major archaeological expedition lead by Richard MacNeish.
Fortunately, the Robert S. Peabody Museum in Andover, MA, took excellent care of the ancient maize specimen–one of the five oldest known in the world–for decades. Wales explains that this particular cob and the DNA within it had been unusually well preserved.
"Archaeological specimens frequently have high levels of bacterial DNA due to decomposition and soil contaminants," he says. "However, during genetic testing of ancient cobs, we were astonished to find that 70 percent of the DNA from the Tehuacan162 cob was from the plant!" Most other ancient samples contain less than 10 percent plant DNA.
Tehuacan162 didn't have hard seed coats like its wild ancestor would have. But, the ancient cob is less than a tenth of the size of modern cobs, at less than two centimeters long. In addition, the ancient cob produced only eight rows of kernels, about half that of modern maize. That led the researchers to suspect that its genes would offer clues on the early stages of maize domestication.
To make the most of the small sample, Wales and Ramos-Madrigal used cutting-edge paleogenomic techniques. They extracted DNA with a method designed to recover ultra-short DNA, taking special care to avoid losing any genetic material. As a result, the researchers were able to prepare sufficient DNA for sequencing while still preserving enough of the sample to determine the cob's precise age via radiocarbon dating.
The new findings offer an informative snapshot in the 10,000-year evolutionary history of maize and its domestication, the researchers say. In addition to elucidating how maize provided a dietary foundation for ancient civilizations like the Maya, such studies can also aid in understanding and improving commercially important lines of modern maize, the researchers say.
"This is only the beginning of the story," Ramos-Madrigal says. "Humans dispersed maize across the Americas very quickly and very successfully. We want to know how humans dispersed it, which routes they took, and how maize adapted to such diverse environments."
###
This research was supported by the Lundbeck Foundation, the Danish Council for Independent Research, and the Danish National Research Foundation .
Current Biology, Ramos-Madrigal et al.; "Genome Sequence of a 5,310-Year-Old Maize Cob Provides Insights into the Early Stages of Maize Domestication" http://www.cell.com/current-biology/fulltext/S0960-9822(16)31120-4
Current Biology (@CurrentBiology), published by Cell Press, is a bimonthly journal that features papers across all areas of biology. Current Biology strives to foster communication across fields of biology, both by publishing important findings of general interest and through highly accessible front matter for non-specialists. Visit: http://www.cell.com/current-biology. To receive Cell Press media alerts, contact [email protected].
Media Contact
Joseph Caputo
[email protected]
617-397-2802
@CellPressNews
http://www.cellpress.com
############
Story Source: Materials provided by Scienmag