HKU chemical biologists identify a new histone mark that regulates chromatin structure during gene expression and DNA repair
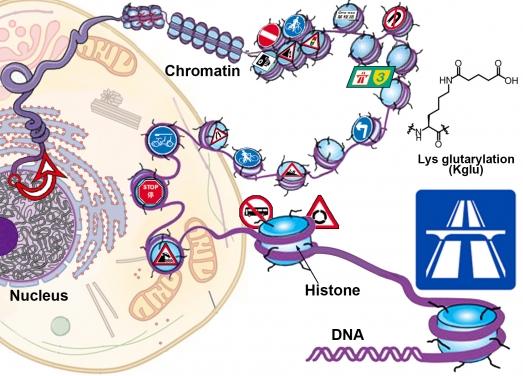
Credit: @The University of Hong Kong
A research team led by Dr Xiang David Li, Associate Professor from the Department of Chemistry, in collaboration with Dr Karen Wing Yee Yuen from the School of Biological Sciences and Dr Jason Wing Hon Wong from the School of Biomedical Sciences at The University of Hong Kong (HKU), revealed a new fundamental mechanism by which a cell can make necessary changes in its chromatin structure in response to different DNA-associated processes such as gene expression and DNA damage repair. The findings were recently published in the prestigious scientific journal Molecular Cell.
Imagine, for a moment, that you are now living in the centre of an ever-shifting maze: paths moving about and breaking apart, wide, straight roads curling up and shrinking into tight, winding tracks, new roads appearing from what used to be dead-ends. Travelling through this labyrinth, the only thing guiding you on the way is various road signs (e.g., “STOP”, “SLOW”, “ONE WAY” and “DO NOT ENTER”) that indicate whether and how you may traverse different paths.
One such maze lies within our every cell: the chromatin, in which DNA is packaged with proteins called histones. Packaging of DNA can be tighter or looser in different regions of chromatin. While a loose packaging indicates an “active” or a gene “ON” region, a tight compaction means a “silent” or a gene “OFF” region. Interestingly, the chromatin also contains various “road signs” in the form of chemical modifications to histones (or histone marks) that indicate the active, inactive or damaged regions of the chromatin and give order to various chromatin-associated machineries in the regulation of gene expression, DNA replication and damage repair. While some well-known chromatin “road signs” such as lysine acetylation (Kac) and methylation (Kme) have been well characterised, the biological meanings of many other “signs”, particularly those newly identified histone marks, remain mysterious.
In a search for new chromatin “road sign”, Dr Li’s team discovered a novel histone mark, lysine glutarylation (Kglu) at histone H4, Lysine-91 (H4K91glu) from human cells. “I was so excited to find this new histone mark. Now imagine that in the maze you come across a new sign that you’ve not seen before. Who puts it there? What does it mean?” said Dr Xiucong Bao, a postdoctoral fellow in Dr Li’s lab and the first author of the study, but she also notes the great challenge to uncover these mysteries, for which she has spent more than five years on the project. The researchers finally found that this mark is especially abundant in promoter regions of active, “open” chromatin where genes are highly expressed – equivalent to a road sign in the maze showing “expressway”. “We believe that H4K91glu is a “sign” for activation of gene expression,” said Dr Li. “And this ‘sign’ seems to be conserved in evolution, as we found it in not only human but also mouse, fly, worm and even baker’s yeast cells.”
Besides marking the active genomic region, H4K91glu in fact directly contributes to the formation of the more open accessible chromatin structure facilitating gene expression. Dr Li’s team found in this study that H4K91glu destabilizes nucleosome, the basic repeating unit of chromatin, and leads to activation or “opening” of chromatin. “It makes perfect sense if you know chemistry, as the mark puts a negative charge on an originally positively charged lysine residue. It therefore causes a charge-charge repulsion within the nucleosome and makes it more prone to falling apart,” says Dr Li.
Much like the ever-shifting maze, chromatin packaging is highly dynamic. A compacted region of chromatin at this moment can quickly change to a relaxed one at the next moment, which allows fast switching between gene ON and OFF states. Meanwhile, when chromatin structure is changed at a specific region, the old “road signs” (i.e., histone marks) are taken off and the new ones are installed by enzymes called histone mark “erasers” and “writers”, respectively. “To understand a histone mark, it is key to find its “writer” and “eraser”, says Li, whose team further identified the enzymes that “write” and “erase'”= the H4K91glu mark: KAT2A, working together with the α-ketoadipate dehydrogenase (α-KADH) complex, adds the H4K91glu mark, while SIRT7 works to remove it. The researchers went on to demonstrate that H4K91glu must be removed by SIRT7 such that the open chromatin region could be inactivated and condensed during cell division or at local DNA damage site.
In summary, Dr Li’s study identified H4K91glu as a novel histone mark and unraveled its regulation and function in chromatin structure and dynamics, putting us one step closer towards deciphering the yet mysterious chromatin maze. The findings from this study also laid the foundation for elucidating how this novel histone mark contributes to human health and disease and will open opportunities for development of therapeutic agents for the treatment of human diseases associated with mis-regulation of histone H4K91glu and chromatin structure.
###
This research is supported by General Research Fund, Collaborative Research Fund and the Area of Excellence Scheme from Hong Kong Research Grants Council, and the National Natural Science Foundation of China.
For more information about the paper “Glutarylation of Histone H4 Lysine 91 Regulates Chromatin Dynamics” at ‘Molecular Cell‘, please visit:
https:/
For more information about Dr Xiang David Li and his research group, please visit their group’s webpage: https:/
Media Contact
Cindy Chan
[email protected]
Original Source
https:/
Related Journal Article
http://dx.




