Tagging produces detailed catalog of transcription factors key to making each cell type
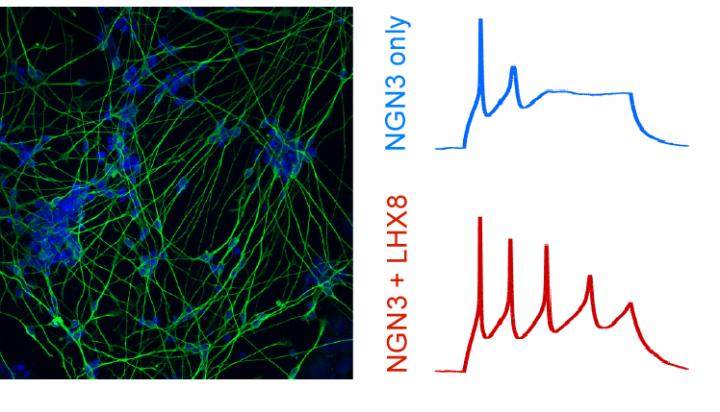
Credit: Gersbach Lab, Duke University
DURHAM, N.C. – A team of biomedical engineers at Duke University has created a new way to turn stem cells into a desired cell type by mastering the language of gene regulatory networks.
Programming stem cells into other cell types is not a new idea. Several methods already exist, but the results have left something to be desired. Often, programmed stem cells do not mature correctly when cultured in the lab, so researchers seeking adult neuron cells for an experiment might end up with embryonic neurons, which won’t be able to model late-onset psychiatric and neurodegenerative conditions.
“The cells might seem right at first glance,” said Josh Black, the Duke Ph.D. student who led the work in Charles Gersbach’s lab, “but they are often missing some of the key properties you want in those cells.”
Using CRISP gene editing, the lab led by Gersbach, The Rooney Family Associate Professor of Biomedical Engineering and the Director of the Center for Advanced Genomic Technologies, created a method to identify which transcription factors – master controllers of gene activity – were essential to making a good neuron.
Their work, appearing Dec. 1 in Cell Reports, demonstrates the potential of the approach to make mature adult neurons, but it could be applied to program any cell type.
CRISPR technology is most often used for editing DNA sequences, known as “genome editing,” in which the Cas9 protein is bound to a guide RNA that directs Cas9 to cut the DNA at a specific location, leading to changes in the DNA sequence. “DNA editing has been widely used to alter gene sequences, but that doesn’t help in situations where the gene is turned off,” Gersbach said.
A deactivated Cas9 (dCas9) protein, though, will attach to the DNA without cutting it. In fact, it typically won’t do anything without another molecule attached or recruited to it. Gersbach and his colleagues have previously reported a variety of methods to attach different molecular domains to the dCas9 protein can that will tell a cell to turn on a gene and remodel chromatin structure.
When Black joined Gersbach’s lab, he was interested in using these tools to turn on genes that could convert one cell type into another to create better disease models.
In 2016, Black and Gersbach reported an approach to use the CRISPR-based gene activators to turn on gene networks that would convert fibroblasts, an easily accessible cell type that makes up connective tissue, to neuronal cells. This study targeted gene networks that were known to be associated with neuronal specification, but did not generate cells with all of the properties needed to make effective disease models. However, the right gene networks to generate those desired cells were unknown, and there are were thousands of possibilities encoded in the human genome. So Black and Gersbach devised a strategy to test all of the networks in a single experiment.
They started with pluripotent stem cells, since this cell type should be able to become any other cell in the human body. To make mature neurons from stem cells, the team engineered stem cells that fluoresced red once they became neuronal. The brighter the fluorescence, the stronger the push towards a neuronal fate. Then they made a pooled library of thousands of guide RNAs targeted to all of the genes that encode transcription factors in the human genome. Transcription factors are the master regulators of gene networks, so to make the desired neurons, they have to get all of the right transcription factors turned on.
They introduced the CRISPR gene activator and guide RNA library into the stem cells so that each cell only received a single guide RNA, and therefore turned on its particular corresponding transcription factor gene target. Then they sorted the cells based on how red they became and sequenced the guide RNAs in the most and least red cells, which told them which genes, when turned on, made the cells more or less neuronal.
When they profiled the gene expression from the stem cells engineered with the guide RNAs, the results suggested that the corresponding cells generated more specific and more mature types of neurons. They also found genes that worked together when targeted simultaneously. Moreover, the experiment revealed factors that antagonized the neuronal commitment of the stem cells, and when they used CRISPR-based repressors of those genes, they could also enhance the neuronal specification.
However, these results were all just measuring markers of neurons. To know if these engineered cells truly recapitulated the function of more mature neurons, they needed to test their ability to transmit electrical signals.
For this, they turned to Professor Scott Soderling, the George Barth Geller Distinguished Professor for Research in Molecular Biology and Chair of the Duke Department of Cell Biology. Shataakshi Dube, a grad student in Soderling’s lab, used a technique known as patch clamp electrophysiology to measure the electrical signals inside the newly formed neurons. By poking a tiny hole in the cell with a very small pipette, she could look inside the neuron and see if it was transmitting electrical signals known as action potentials. If so, the team knew the neuronal cell had properly matured. In fact, the neurons engineered to activate a particular pair of transcription factor genes were more functionally mature, emitting more action potentials more frequently.
“I was curious but skeptical on how neuronal these stem cells could become,” Dube said, “but it was remarkable to see how much these programmed cells looked just like normal neurons.”
The process from stem cell to mature neuronal cell took seven days, dramatically shortening the timeframe compared to other methods that take weeks or months. This faster timeline has the potential to significantly accelerate the development and testing of new therapies for neurological disorders.
Creating better cells will help researchers in a number of ways. Diseases like Alzheimer’s disease, Parkinson’s disease, and schizophrenia most often occur in adults and are difficult to study because making the right cells in the lab is challenging. This new method can allow researchers to better model these diseases and others. It can also help with drug screening, as different cells respond to drugs differently.
More broadly, the same method for screening transcription factor genes and gene networks could be used to improve methods to make any cell type, which could be transformative for regenerative medicine and cell therapy.
For example, Gersbach’s group reported a method for using CRISPR-based gene activation to convert human stem cells into muscle progenitor cells that could regenerate damaged skeletal muscle tissue earlier this year.
“The key to this work is developing methods to use the power and scalability of CRISPR-based DNA targeting to program any function into any cell type,” Gersbach said. “By leveraging the gene networks already encoded in our genome, our control over cell biology is dramatically improved.”
###
CITATION: “Master Regulators and Cofactors of Human Neuronal Cell Fate Specification Identified by CRISPR Gene Activation Screens,” Joshua B Black, Sean R McCutcheon, Shataakshi Dube, Alejandro Barrera, Tyler S Klann, Grayson A Rice, Shaunak S Adkar, Scott H Soderling, Timothy E. Reddy and Charles A Gersbach. Cell Reports, Dec. 1, 2020, DOI: 10.1016/j.celrep.2020.108460
Media Contact
Karl Bates
[email protected]
Related Journal Article
http://dx.




