Credit: ©Science China Press
Soybean (Glycine max [L.] Merr.) is one of the most important crops, providing more than half of global oilseed production and more than a quarter of the world's protein for food and animal feed. Studies have indicated that the cultivated soybean was domesticated in China approximately 5,000 years ago and then disseminated worldwide. During the introduction and dissemination process, soybean has undergone strictly genetic bottlenecks, resulting in the accessions from different geographic areas possibly exhibiting high genetic diversity. The current soybean reference genome was sequenced from Williams 82, which is a cultivar domesticated in America. Asia is one of the largest soybean planting and consuming areas, its soybean production is essential for global food security. A high-quality reference genome is crucial for functional analysis of a species. Therefore, it is necessary to assemble a new high-quality soybean genome from Asian soybean accessions to facilitate Asia soybean functional genomics study and elite cultivar improvement.
Biologists from the Institute of Genetics and Developmental Biology, Chinese Academy of Science, University of Science and Technology of China, Jiangsu Academy of Agricultural Sciences, and Berry Genomics Corporation assembled a new soybean genome from a Chinese soybean accession "Zhonghuang 13" by combination of SMRT, Hi-C and optical mapping data.
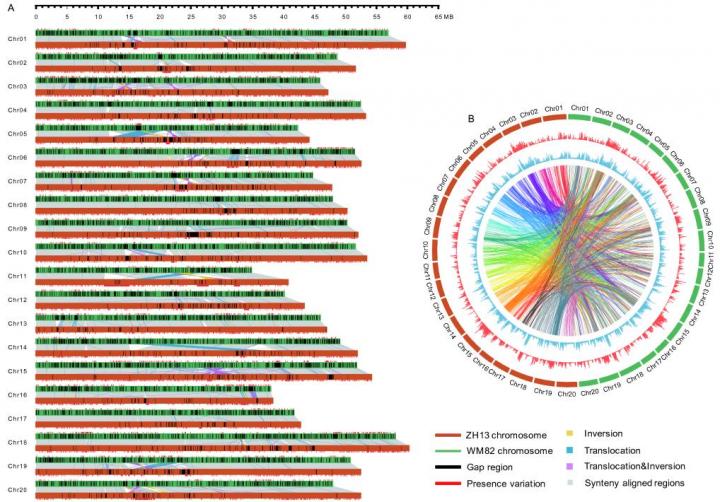
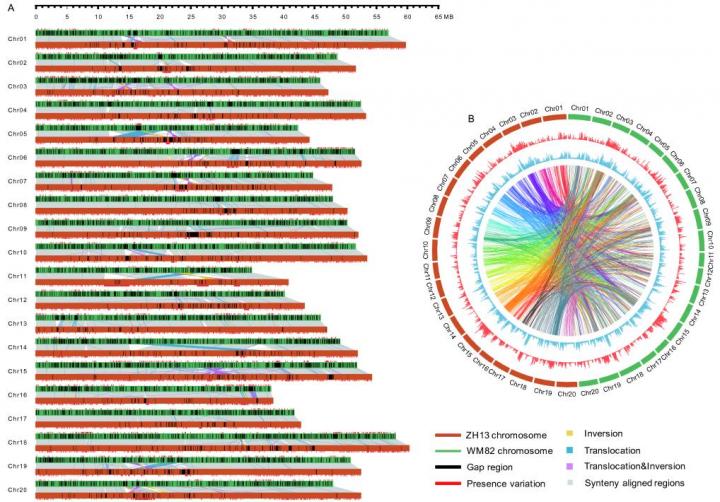
"Our finally assembled genome is 1.025 Gb. Its contig N50 is 3.46 Mb, and scaffold N50 is 51.87 Mb. To the best of our knowledge, it is the fourth contiguous plant genome that has been reported to date." said Dr. Yanting Shen, the senior author for this work. "Comparing to the previously commonly used soybean reference genome, the quality of our new assembled genome is significantly improved, it has longer total sequence length, higher contig N50 and scaffold N50, and fewer gaps". Moreover, after careful comparison of the genome sequences between Zhonghuang 13 and Williams 82, the research team found that the Zhonghuang 13 genome displays large number of genetic variations with that of Williams 82. "We identified more than 250,000 structure variation events, including 1,404 translocations, 161 inversions, 1,233 translocation & inversions, 505,506 indels (1-99 bp) and 17,409 accession specific insertions (>=100 bp)", said Yanting.
"This work is a great team work that collaborated by the scientists from different areas. For example, to make the new genome annotation more accurate and informative, we asked help from Dr. Jianchang Du, an expert in transposable elements study" said the corresponding author Dr. Zhixi Tian, a scientist working on soybean functional genome. Through comprehensive analyses, a total of 36,429 transposable elements and 52,051 protein coding genes were annotated in the new genome. "QTL and GWAS are useful approaches to investigate the loci controlling traits. However, it is difficult to identify the causal gene directly from QTL and GWAS because they usually result in large candidate regions. Gene co-expression network is a powerful approach to explore gene regulatory relationships and to predict gene function. We think a gene co-expression network may assist important agronomic genes mining when combing with QTL and GWAS. Therefore, we asked help from Dr. Shisong Ma, an expert on gene co-expression network from University of Science and Technology of China", said Zhixi.
"To test this hypothesis, we established a comprehensive gene co-expression network for the annotation genes using transcriptome datasets from 1,978 soybean RNA-seq that deposited in the NCBI Sequence Read Archive (SRA). This network contain 39,967 genes and 330,864 co-expressed gene pairs", said Dr. Shisong Ma, the collaborator and one of the co-corresponding author. The usage of gene co-expression network was tested by exploring new genes related to soybean flowering time and linoleic acid content. "Our gene co-expression network narrowed number of soybean flowering time candidate genes from 7,971 to 26 directly, which was effective in candidate gene mining," said Yanting, "In further, we confirmed the function for one candidate gene SoyZH13_16G177400 by correlating its haplotype and phenotype information in our previously re-sequenced nature population." Besides the gene related to flowering time, another gene controlling soybean linoleic acid content also be confirmed using the same strategy. Dr. Baoge Zhu, a biologist that has been working on soybean breeder for more than twenty years from the Institute of Genetics and Developmental Biology, Chinese Academy of Science, commented "This method is very novel and helpful for our soybean researchers and breeders. We can use it to dig candidate genes effectively for agronomic important phenotypes."
All the authors hope that this new genome will facilitate legume genomics research and soybean crop improvement in the future.
###
This work was supported by the National Natural Science Foundation of China (91531304, 31525018, 31370266, 31788103), the "Strategic Priority Research Program" of the Chinese Academy of Sciences (XDA08000000), and the State Key Laboratory of Plant Cell and Chromosome Engineering (PCCE-KF-2017-03).
See the article:
Shen, Y., Liu, J., Geng, H., Zhang, J., Liu, Y., Zhang, H., Xing, S., Du, J., Ma, S., and Tian, Z. (2018). De novo assembly of a Chinese soybean genome. Sci China Life Sci 61, https://doi.org/10.1007/s11427-018-9360-0
http://engine.scichina.com/publisher/scp/journal/SCLS/doi/10.1007/s11427-018-9360-0
Media Contact
Tian Zhixi
[email protected]
http://www.scichina.com/
Related Journal Article
http://dx.doi.org/10.1007/s11427-018-9360-0