Credit: Pavel Odinev / Skoltech
A team of Skoltech researchers from the Severinov laboratory and their colleagues have identified the way in which a component of a two-part bacterial self-defense system from the toxin-antitoxin family works, leading to cell dormancy that helps fight off bacterial viruses, antibiotics and other insults.
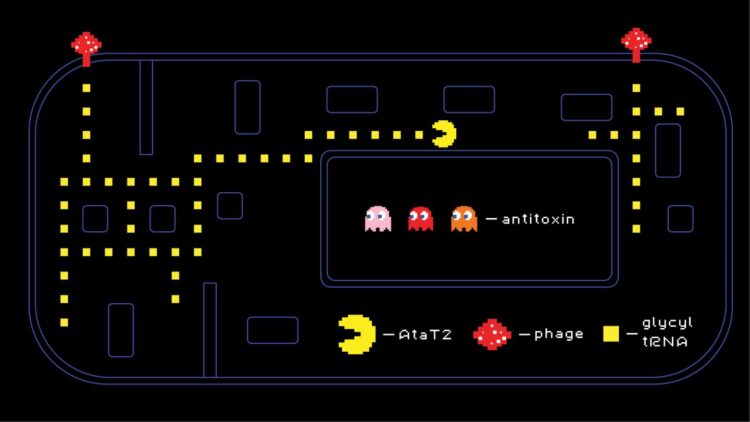
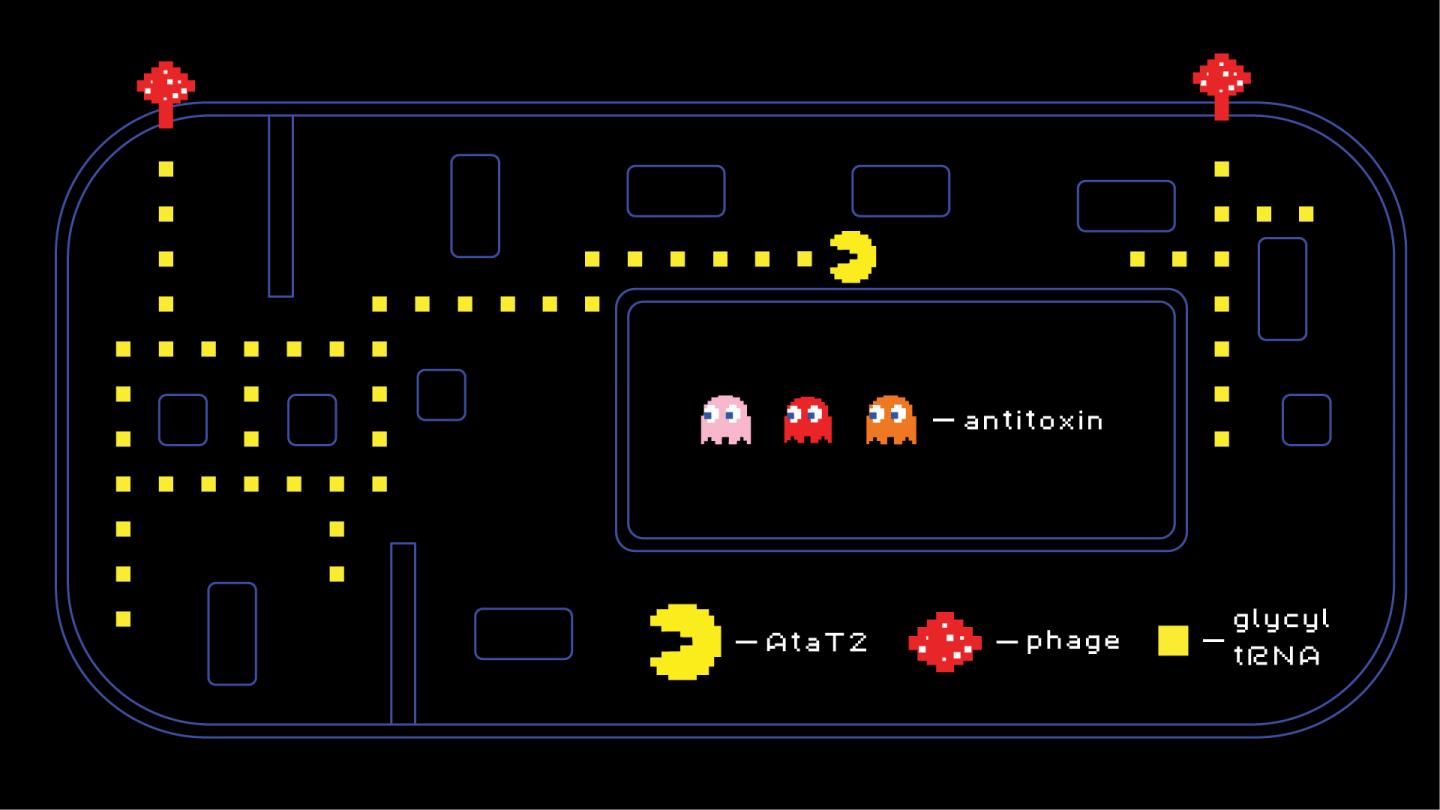
Toxin-antitoxin systems are a class of multipurpose mechanisms that bacteria can use, among other things, against phage infections. Two adjacent genes encode two proteins, a toxin disrupting various cellular processes and an antitoxin that inhibits the toxin’s activity. Much like Fluffy the three-headed dog from the Harry Potter series, the toxin is “dormant” while the music is playing (i.e. while the antitoxin is present), but under stress conditions — say, when a bacteriophage attacks — the antitoxin is no longer produced and the toxin is “unleashed”, disrupting protein synthesis needed for viral replication.
“Toxin-antitoxin systems are very widespread in bacteria, and many people tried to answer what their raison d’être is, why they exist. The answer appears to be elusive, and the story of assigning biological function to these systems is full of drama, retraction of papers and the like. The answer may be the childish “just because”: they may be selfish elements concerned with their own propagation more than with the well-being of the cell, which, however, does not make these systems less interesting to study or reduce the knowledge about their function to practice,” says Konstantin Severinov, Skoltech professor and a coauthor of the paper.
There are various types of toxin-antitoxin systems, which are classified depending on how exactly the antitoxin blocks the toxin. Though thousands of these pairs have been predicted with the help of bioinformatics, only a handful has been thoroughly characterized. Many toxins are ribonucleases that degrade RNA, but some have different activities.
A team of researchers led by Severinov and Svetlana Dubiley of Skoltech Centre for Life Sciences studied AtaT2, a representative of a rare class of toxins called GNAT (for Gcn5-related N-acetyltransferase). They show that this toxin disrupts translation, or the synthesis of proteins by the ribosome, by targeting transfer RNAs for glycine, a common protein-building amino acid.
The scientists modified E. coli to express the toxin-antitoxin system genes on demand and then conducted in vivo tests to determine how the toxin works by observing the behavior of intoxicated cells. They also performed in vitro analysis and, by combining the two, found that the toxin interferes with translation by stalling ribosomes at the glycine codons in the protein encoding sequence, so that most of them are unable to complete the process and build a protein.
Interestingly, the antitoxin counterpart of AtaT2 does not have any glycyl residues for the toxin to target, so its synthesis is unaffected by the toxin. The researchers speculate that this might be a built-in feedback loop preventing too much AtaT2 from being produced and helping cells recover from its toxic action.
If the hypothesis about the essentially selfish nature of toxin-antitoxin systems is correct, one can imagine that different systems compete with each other for their hosts, the bacteria they inhabit. If so, the targets of related toxins should diverge with time. The Severinov team hypothesizes that GNAT toxins may thus diversify to target transfer RNAs specific for each of the 20 genetically encoded amino acids.
“If that is true, a panel of such toxins, whatever their biological function may be, can provide a powerful tool to control each of the elemental steps of protein synthesis inside the cell and may lead to development of powerful new antibiotics,” Severinov notes.
###
Other organizations involved in this research include the RAS Institute of Gene Biology, Lomonosov Moscow State University, University of Illinois, Rowan University School of Osteopathic Medicine, and Waksman Institute for Microbiology.
Media Contact
Alina Chernova
[email protected]
Related Journal Article
http://dx.