St. Jude Children’s Research Hospital scientists offer insight into the toxicity at the heart of devastating neurological disorders
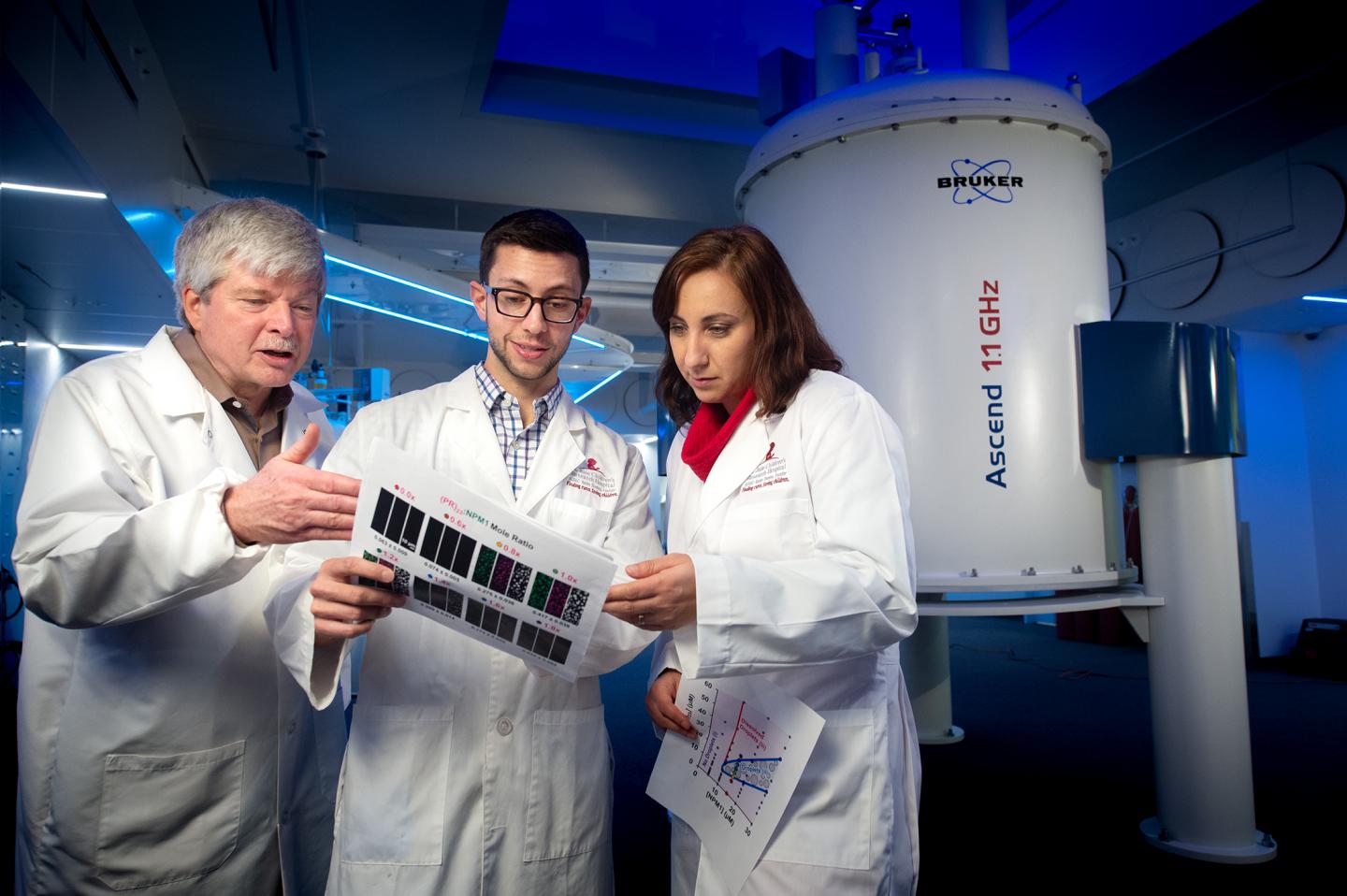

Credit: St. Jude Children’s Research Hospital / Seth Dixon
St. Jude Children’s Research Hospital scientists have cracked the mystery surrounding the most common genetic cause of amyotrophic lateral sclerosis (ALS), or Lou Gehrig’s disease. The research suggests possible new approaches to diagnosis and treatment of the lethal disorder. The findings appear online today in the journal Molecular Cell.
About 35 percent of ALS cases are associated with mutations in the gene C9orf72, making it the most common genetic cause of ALS and another neurological disease, frontotemporal dementia (FTD). The mutation leads to a dramatic increase in the number of short repetitive DNA sequences and results in the formation of abnormal repetitive proteins of varying lengths. These proteins are referred to as dipeptide repeat polypeptides (DPRs) Two of the formed DPRs contain the amino acid arginine, and are particularly toxic to neurons.
Until now, key details about the molecular mechanisms involved were uncertain.
“We have identified the protein, nucleophosmin, as a site of DPR toxicity,” said corresponding author Richard Kriwacki, Ph.D., a member of the St. Jude Department of Structural Biology. “We also show that DPR toxicity is exquisitely length dependent. In the future, DPR length may have prognostic value for people with a diagnosis of ALS.”
ALS, DPRs and disease
ALS is a rapidly progressive neurological disease that kills nerve cells responsible for voluntary muscle control. Frontotemporal dementia is caused by the loss of nerve cells in the brain. Currently there is no cure for either disease.
Typically, a segment of the C9orf72 gene is repeated 20 to 30 times or less. However, those with ALS and FTD have hundreds or even thousands of the repeats, which then cause the formation of DPRs. Previous research from Kriwacki and others reported that the toxic (arginine-containing) DPRs disrupt assembly and function of the nucleolus, the largest membrane-less organelle in cells. This study highlights how they disrupt nucleolar assembly. The study also shows that longer DPRs are dramatically more toxic to cells.
The nucleolus resides in the nucleus and is where the protein factories of cells (called ribosomes) are assembled. Unlike the nucleus, the nucleolus lacks a membrane. Membrane-less organelles like the nucleolus rely on a process called liquid-liquid phase separation to form and give cells flexibility to respond to changing conditions. The same process explains why oil forms droplets in water.
Uninvited guests
Kriwacki and his colleagues showed the toxic DPRs disrupt cell function by tightly binding to key regions of nucleophosmin, displacing other binding partners that help maintain the nucleolus and ribosome assembly. The greater the concentration of toxic DPRs, the faster the membrane-less nucleolus is altered and dissolved.
DPRs bind and sequester nucleophosmin within large complexes, causing the nucleolus to partially dissolve. Researchers showed the toxic DPRs also disrupted cell function by binding and isolating a key component of ribosomes–ribosomal RNAs.
NMR
A 1.1 GHz nuclear magnetic resonance (NMR) system scheduled for installation at St. Jude later this year solidified the findings, Kriwacki said. The device, which is currently at the manufacturer in Zürich, Switzerland, boasts the world’s strongest high-resolution NMR magnet to enhance the study of biological samples. Researchers used it in this study to reveal in greater detail how toxic DPRs bound to intrinsically disordered regions of nucleophosmin.
Clinical possibilities
“The work also provides a new direction for thinking about possible therapies to target toxic DPRs and their sites of action in patient cells,” said first author Michael White, Ph.D., a postdoctoral fellow in Kriwacki’s laboratory.
“Success will take totally innovative approaches,” Kriwacki said. “But at least now we know what needs to be targeted.”
###
The other authors are Diana Mitrea, Peipei Zhang, Devon Cassidy, Amanda Nourse, Aaron Phillips, and Michele Tolbert, all of St. Jude; J. Paul Taylor, of St. Jude and the Howard Hughes Medical Institute; and Christopher Stanley, of Oak Ridge National Laboratory, Oak Ridge, Tennessee.
The study was funded in part by grants (GM115634, NS097974, CA23944) from the National Institutes of Health; the Howard Hughes Medical Institute, the St. Jude Research Collaborative on Membrane-less Organelles and ALSAC, the fundraising and awareness organization of St. Jude.
Media Contact
Corey Carmichael
[email protected]
Original Source
http://www.
Related Journal Article
http://dx.