New computational tools developed at Stony Brook University help characterize protein structures and identify new treatments for COVID-19
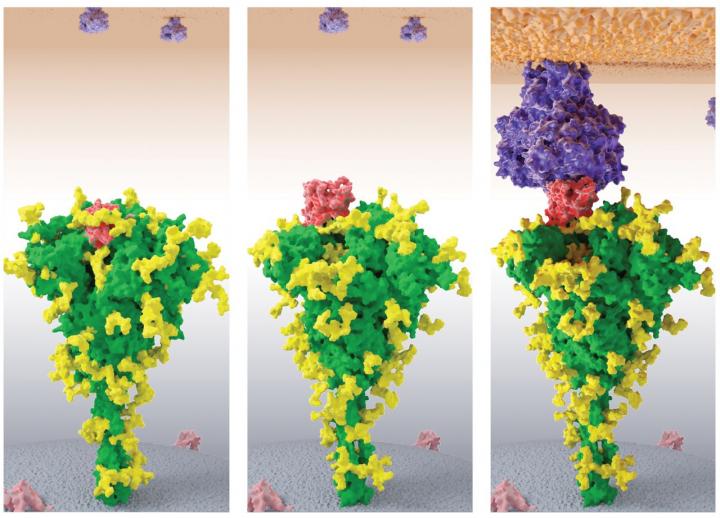
Credit: Lucy Fallon, Laufer Center, Stony Brook University
In the last five decades, we’ve learned a lot about the secret lives of proteins — how they work, what they interact with, the machinery that makes them function — and the pace of discovery is accelerating.
The first three-dimensional protein structure began emerging in the 1970s. Today, the Protein Data Bank, a worldwide repository of information about the 3D structures of large biological molecules, has information about hundreds of thousands of proteins. Just this week, the company DeepMind shocked the protein structure world with its accurate, AI-driven predictions.
But the 3D structure is often not enough to truly understand what a protein is up to, explains Ken Dill, director of the Laufer Center for Physical and Quantitative Biology at Stony Brook University and a member of the National Academy of Sciences. “It’s like somebody asking how an automobile works, and a mechanic opening the hood of a car and saying, ‘see, there’s the engine, that’s how it works.'”
In the intervening decades, computer simulations have built upon and added to the understanding of protein behavior by setting these 3D molecular machines in motion. Analyzing their energy landscapes, interactions, and dynamics has taught us even more about these prime movers of life.
“We’re really trying to ask the question: how does it work? Not just, how does it look?” Dill said. “That’s the essence of why you want to know protein structures in the first place, and one of the biggest applications of this is for drug discovery.”
Writing in Science magazine in November 2020, Dill and his Stony Brook colleagues Carlos Simmerling and Emiliano Brini shared their perspectives on the evolution of the field.
“Computational Molecular Physics is an increasingly powerful tool for telling the stories of protein molecule actions,” they wrote. “Systematic improvements in forcefields, enhanced sampling methods, and accelerators have enabled [computational molecular physics] to reach timescales of important biological actions…. At this rate, in the next quarter century, we’ll be telling stories of protein molecules over the whole lifespan, tens of minutes, of a bacterial cell.”
SPEEDING SIMULATIONS
Decades after the first dynamic models of proteins, however, computational biophysicists still face major challenges. To be useful, simulations need to be accurate; and to be accurate, simulation needs to progress atom by atom and femtosecond (10^-12 seconds) by femtosecond. To match the timescales that matter, simulations must extend over microseconds or milliseconds — that is, millions of time-steps.
“Computational molecular physics has developed at a fast clip relatively speaking, but not enough to get us into the time and size and motion range we need to see,” he said.
One of the main methods researchers use to understand proteins in this way is called molecular dynamics. Since 2015, with support from the National Institutes of Health and the National Science Foundation, Dill and his team have been working to speed up molecular dynamics simulations. Their method, called MELD, accelerates the process by providing vague but important information about the system being studied.
Dill likens the method to a treasure hunt. Instead of asking someone to find a treasure that could be anywhere, they provide a map with clues, saying: ‘it’s either near Chicago or Idaho.’ In the case of actual proteins, that might mean telling the simulation that one part of a chain of amino acids is near another part of the chain. This narrowing of the search field can speed up simulations significantly — sometimes more than 1000-times faster — enabling novel studies and providing new insights.
PROTEIN STRUCTURE PREDICTIONS FOR COVID-19
One of the most important uses of biophysical modeling in our daily lives is drug discovery and development. 3D models of viruses or bacteria help identify weak spots in their defenses, and molecular dynamics simulations determine what small molecules may bind to those attackers and gum up their works without having to test every possibility in the lab.
Dill’s Laufer Center team is involved in a number of efforts to find drugs and treatments for COVID-19, with support from the White House-organized COVID-19 HPC Consortium, an effort among Federal government, industry, and academic leaders to provide access to the world’s most powerful high-performance computing resources in support of COVID-19 research.
“Everyone dropped other things to work on COVID-19,” Dill recalled.
The first step the team took was to use MELD to determine the 3D shape of the coronavirus’ unknown proteins. Only three of the 29 of the virus’ proteins have been definitively resolved so far. “Most structures are not known, which is not a good beginning for drug discovery,” he said. “Can we predict structures that are not known? That’s the primary thing that we used Frontera for.”
The Frontera supercomputer at the Texas Advanced Computing Center (TACC) — the fastest at any university in the world — allowed Dill and his team to make structure predictions for 19 additional proteins. Each of these could serve as an avenue for new drug developments. They have made their structure predictions publicly available and are working with teams to experimentally test their accuracy.
While it seems like the vaccine race is already close to declaring a winner, the first round of vaccines, drugs, and treatments are only the starting point for a recovery. As with HIV, it is likely that the first drugs developed will not work on all people, or will be surpassed by more effective ones with fewer side-effects in the future.
Dill and his Laufer Center team are playing the long game, hoping to find targets and mechanisms that are more promising than those already being developed.
REPURPOSING DRUGS AND EXPLORING NEW APPROACHES
A second project by the Laufer Center group uses Frontera to scan millions of commercially available small molecules for efficacy against COVID-19, in collaboration with Dima Kozakov’s group at Stony Brook University.
“By focusing on the repurposing of commercially available molecules it’s possible, in principle, to shorten the time it takes to find a new drug,” he said. “Kozakov’s group has the ability to quickly screen thousands of molecules to identify the best hundred ones. We use our physics modeling to filter this pool of candidates even further, narrowing the options experimentalists need to test.”
A third project is studying an interesting cellular protein known as PROTAC that directs the “trash collector proteins” of human cells to pick up specific target proteins that they would not usually remove.
“Our cell has smart ways to identify proteins that needs to be destroyed. It gets next to it, puts a sticker on it, and the proteins who collect trash take it away,” he explained. “Initially PROTAC molecules have been used to target cancer related proteins. Now there is a push to transfer this concept to target SARS-CoV-2 proteins.”
Collaborating with Stony Brook chemist Peter Tonge, they are working to simulate the interaction of novel PROTACS with the COVID-19 virus. “These are some of our most ambitious simulations, both in term of the size of the systems we are tackling and in terms of the chemical complexity,” he said. “Frontera is a crucial resource to give us sufficient turnaround times. For one simulation we need 30 GPUs and four to five days of continuous calculations.”
The team is developing and testing their protocols on a non-COVID test system to benchmark their predictions. Once they settle on a protocol, they will apply this design procedure to COVID systems.
Every protein has a story to tell and Dill, Brini and their collaborators are building and applying the tools that help elucidate these stories. “There are some problems in protein science where we believe the real challenge is getting the physics and math right,” Dill concluded. “We’re testing that hypothesis on COVID-19.”
###
Media Contact
Aaron Dubrow
[email protected]
Original Source
https:/
Related Journal Article
http://dx.