Credit: Courtesy of Shouling Xu and Zhiyong Wang.
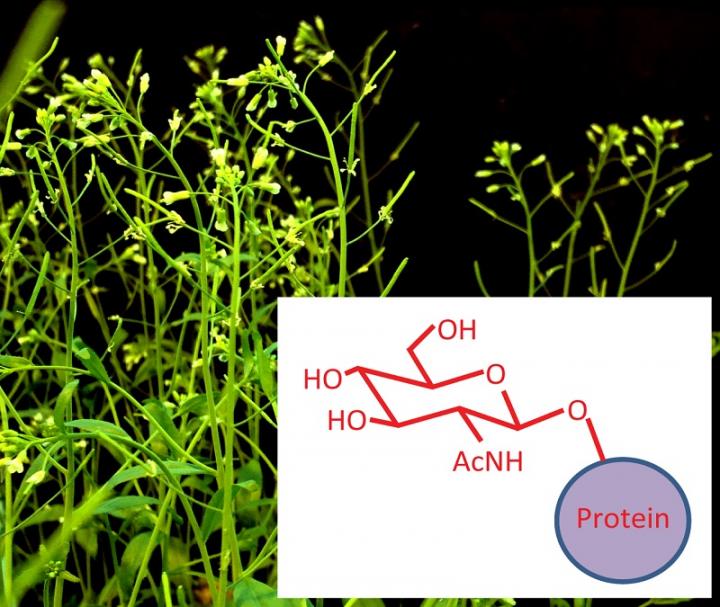
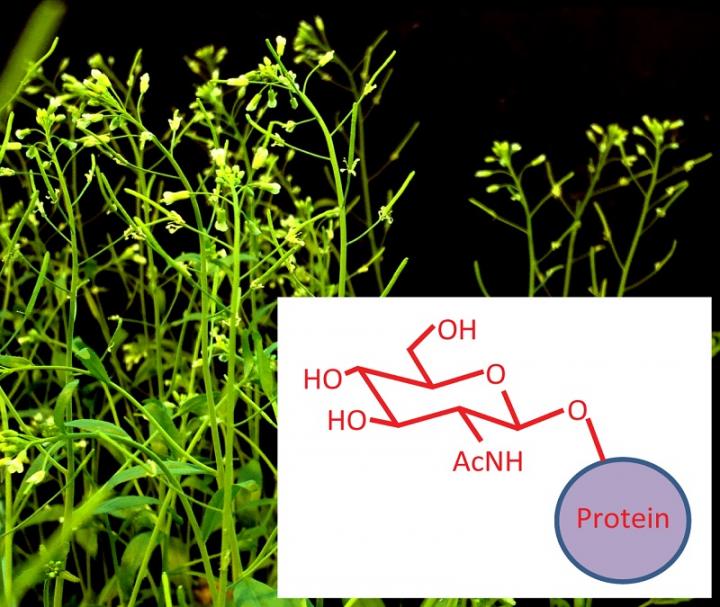
Stanford, CA — New work from Carnegie's Shouling Xu and Zhiyong Wang reveals that the process of synthesizing many important master proteins in plants involves extensive modification, or "tagging" by sugars after the protein is assembled. Their work uncovers both similarity and distinction between plants and animals in their use of this protein modification. It is published by Proceedings of the National Academy of Sciences.
The blueprint for making all proteins is encoded in DNA. The genetic code tells the cellular protein-making apparatus the correct order in which to string together the amino acids that are the building blocks of every protein. Often, after their DNA code has been translated into the amino acid chain, newly synthesized proteins are further modified with different chemical moieties.
A common form of post-translational modification in animal cells has the tongue-twisting name of O-GlcNAcylation (pronounced oh-gluck-nakel-ation). It is a process by which certain amino acids in proteins get attached to a sugar molecule, and this modification impacts a wide array of cellular functions. In animals, changes to this process are associated with neurodegeneration, diabetes, cardiovascular diseases, and cancer. Embryos lacking the enzyme that accomplishes this modification cannot survive.
It was already known that O-GlcNAcylation takes place in plant cells, too. Plants in which the process is impeded show defects in light response, flower development, root growth, and leaf structure, among other things. However, much about which actual plant proteins undergo this modification had remained mysterious until now.
Xu and Wang collaborated with scientists of UCSF and China in identifying plant proteins that undergo this post-production, sugar-adding modification. A number of these affected proteins have regulatory functions that perform the same or similar functions in plants and animals. Many also perform developmental and physiological tasks that are unique to plants, such as flower development and responses to specific plant hormones.
"We identified 262 proteins that are modified by this process," Xu explained. "Furthermore our study provides the framework for a network of O-GlcNAcylation modification in plants, which will help us understand how the functions of these various proteins–and hence the growth processes they control–are regulated according to the availability of sugar in the cells."
"This is a big breakthrough for plant science," said Carnegie Plant Biology Director Sue Rhee. "Shouling and Zhiyong and their colleagues have opened up a whole new area of exploration that will advance our understanding of plant cellular biology."
###
This study was funded by the Chemical Sciences, Geosciences, and Biosciences Division of the Office of Basic Energy Science, Office of Science at the U.S. Department of Energy, the National Institutes of Health, and the Howard Hughes Medical Institute.
The Carnegie Institution for Science (carnegiescience.edu) is a private, nonprofit organization headquartered in Washington, D.C., with six research departments throughout the U.S. Since its founding in 1902, the Carnegie Institution has been a pioneering force in basic scientific research. Carnegie scientists are leaders in plant biology, developmental biology, astronomy, materials science, global ecology, and Earth and planetary science.
Media Contact
Shouling Xu
[email protected]
@carnegiescience
https://carnegiescience.edu/
############
Story Source: Materials provided by Scienmag