Credit: Hasindu Gamaarachchi
The ability to read the genome – all the DNA of an organism – has vast potential to understand human health and disease.
Researchers at the Garvan Institute of Medical Research and UNSW Sydney have published a method to take genome analysis ‘offline’, by adapting a computer algorithm that can perform accurate analysis – with far less computer memory than current programs. The scientists’ algorithm may make it possible to identify infectious diseases in remote locations, or at the hospital bedside, using the computational memory of devices as small as a smartphone.
They published their findings in Scientific Reports on 13 March, 2019.
Genomics without borders
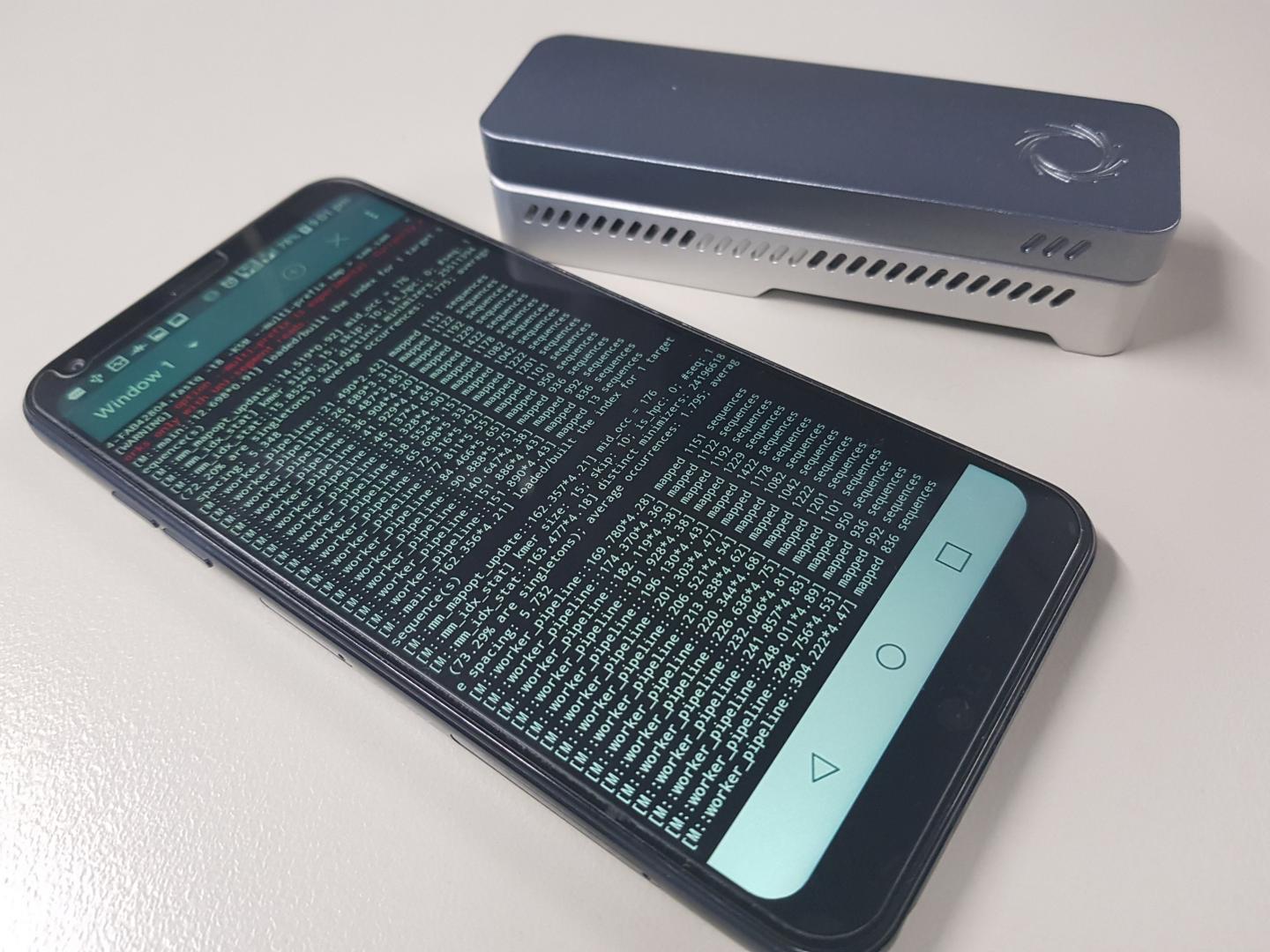
Devices that can sequence entire genomes, such as the Oxford Nanopore Technologies MinION sequencers, are small enough today to clip onto a smartphone – and have already been used to track the Ebola virus in New Guinea and the Zika virus in Brazil.
Such devices are able to create over a terabyte of data in 48 hours, but their use has been limited, because comparing or ‘aligning’ the DNA from an unknown sample to a reference database of known genomes is computationally intensive. Until now, this process was only possible with either high performance computer workstations or an internet connection.
Now, Dr Martin Smith, Team Leader of Genomic Technologies at the Garvan Institute’s Kinghorn Centre for Clinical Genomics, and his team have published a computational method for how to reduce the amount of memory necessary to align genomic sequences from 16GB to 2GB, making it possible for analysis to be done on-the-spot, using the memory available in a typical smartphone.
“We’re focused on making genomic technologies more accessible to improve human health. They’re becoming smaller, but still need to function in remote areas, so we created a method that can analyse genomic data, in real time, on just a mobile device,” explains Dr Smith.
Divide and conquer
The team adapted the Minimap2 program, which aligns DNA sequencing ‘reads’ to a reference library of known genomes. The reference library is usually sorted, or indexed, which helps quickly map the sequencing reads to their corresponding positions in a reference genome.
“The challenge, so far, has been that the reference index requires too much computer memory,” explains Dr Smith. “We took the approach of splitting the reference library up into smaller segments, against which we mapped the DNA reads. Once we finished mapping to the smaller segments, we pool results together and tease out the noise, much like creating a panorama by stitching together smaller photos.”
“Other algorithms, which take a similar approach of splitting up the reference data, produce a lot of spurious and duplicate mappings – just like overlapping photos in the panorama. What we did in this study was fine-tune parameters and select the best mappings across several small indexes. This approach gave us similar accuracy as current standard genomic analyses, which previously required the memory available in high performance computers,” says Dr Smith.
Dr Smith’s team compared the accuracy of their algorithm to standard genomics workflows. Not only did their results reproduce 99.98% of the alignments, but by using the smaller index segments the team could map an additional 1% of sequencing reads.
Dr Smith is optimistic about his technology. “The potential of lightweight, portable genomic analysis is vast – we hope that this technology will one day be applied in the context of point-of-care microbial infections in remote regions, or in doctors’ hands at the hospital bedside,” says Dr Smith.
###
Media Contact
Viviane Richter
[email protected]