Credit: Image courtesy of MIPT Press Office
Biologists and mathematicians from MIPT, Stony Brook University and other scientific research centres have taught a computer to predict the structure of protein complexes in a cell 10 times faster than before. The study has been published in Proceedings of the National Academy of Sciences of the USA.
"The new method enables us to model the interaction of proteins at genome level. This will give us a better understanding of how our cells function and may enable drug development for diseases caused by "incorrect" protein interactions," says Dima Kozakov, a professor at Stony Brook and adjunct professor at MIPT.
A matter of skill
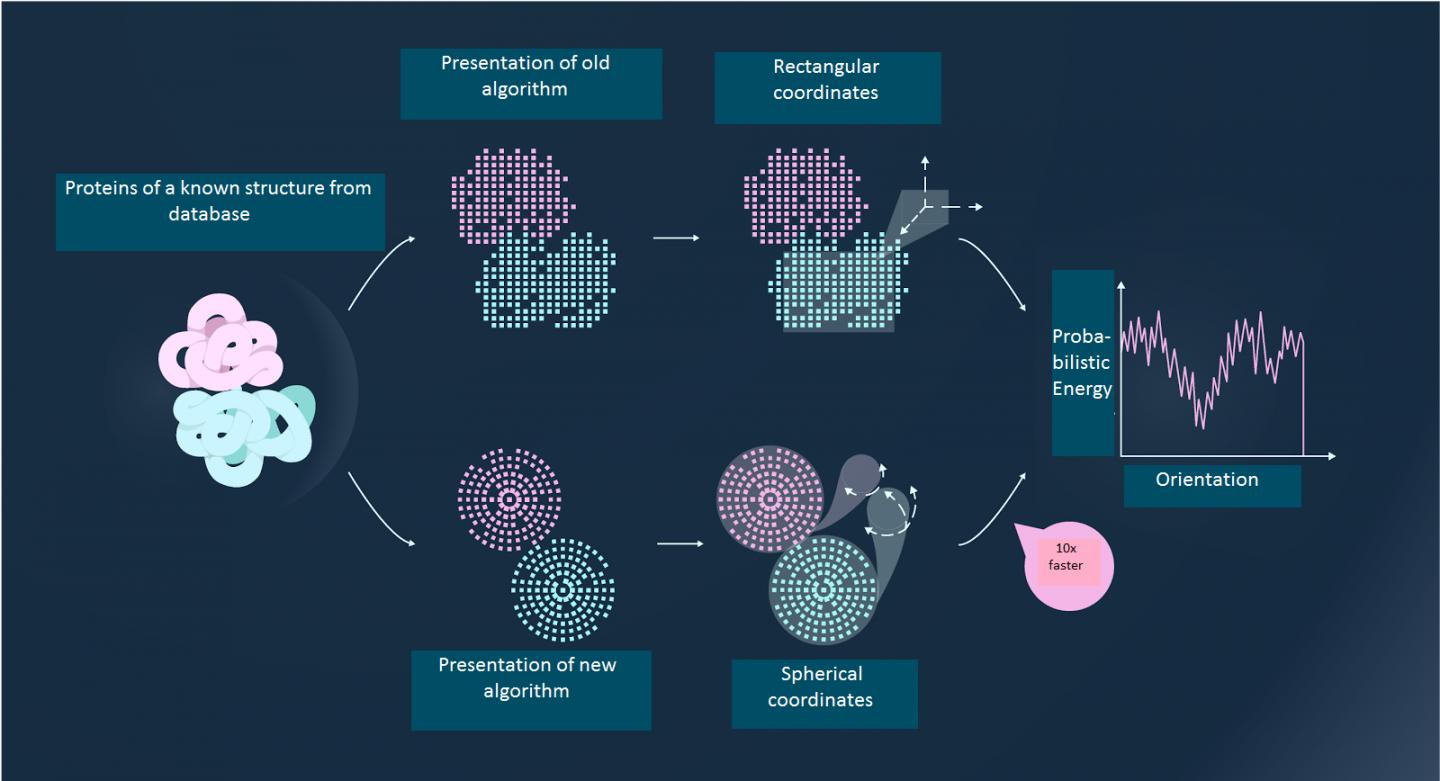
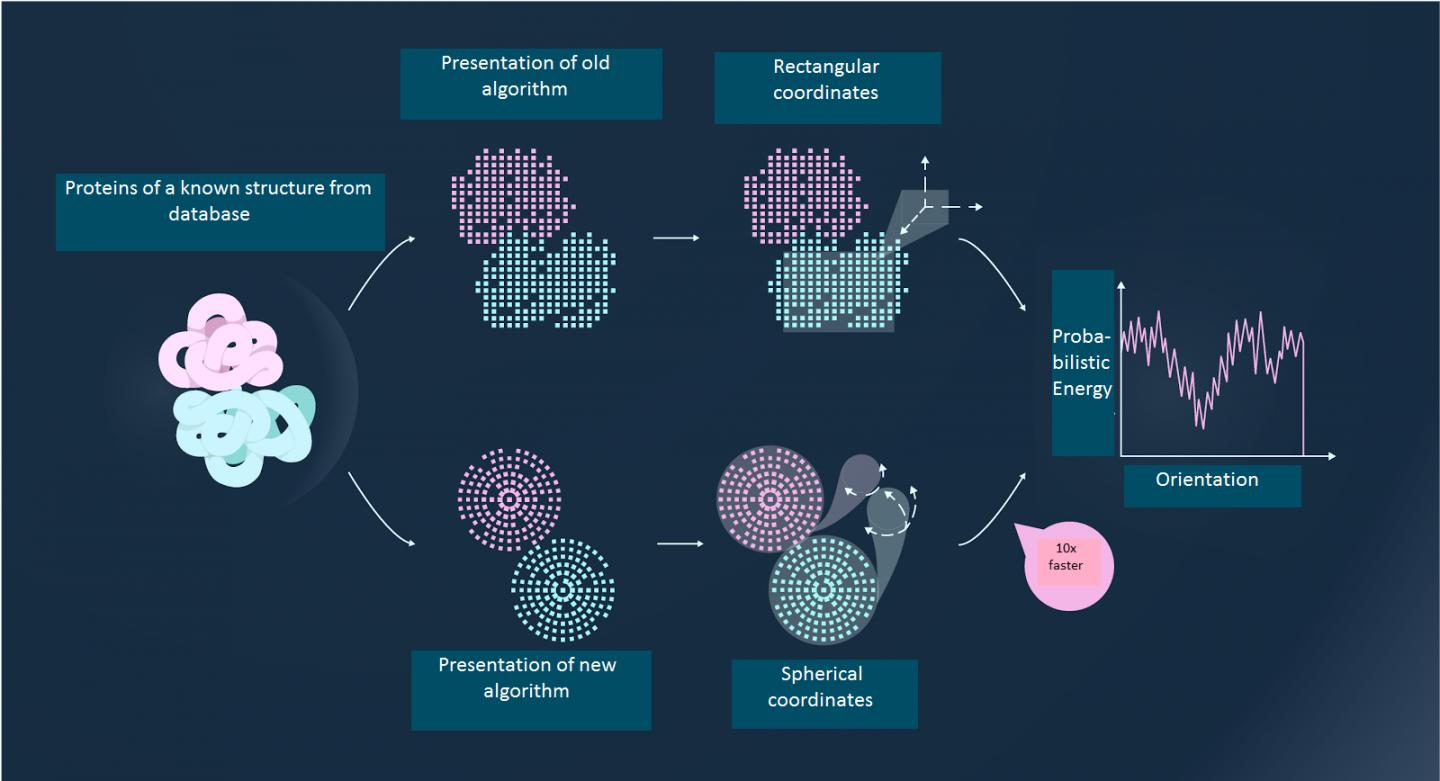
From all the possible alternatives of the orientation of two large molecules in relation to one another, in order to find the one that actually exists scientists solve the following problem: two proteins with a known structure are given. A prediction needs to be made of how they will look when "docked". This is known as the rigid docking method, where the structure of the elements is given and they need to be assembled in the best configuration. In scientific terms, this task is called protein-protein docking.
At first glance, this task appears to be simple and straightforward: assembling the structure of proteins is a matter of skill, just the same as putting together a toy construction set. But, according to the scientists, the computational complexity of such an operation is comparable to assembling all the possible pairs of 10,000 blocks of Lego.
The idea the researchers had was to present proteins as a combination of "quantum surfaces" – certain blocks described by the mathematical tool of quantum mechanics. Using this approach, it is possible to simultaneously calculate the interaction between multiple pairs of protein clusters, rather than examining each pair independently. The new method is up to 100 times faster than the best methods used previously, and it is still accurate. According to the scientists, the program takes 15 minutes to run on a personal computer and is a good alternative to experimental methods of determining protein interactions.
The new algorithm will soon become part of ClusPro – a popular automated system for calculating protein-protein interactions. This resource, which was developed earlier by the authors of the paper, now has more than 15,000 users worldwide. In the latest CAPRI round (Critical Assessment of PRediction of Interactions – a communitywide experiment to determine the structure of proteins), the ClusPro server was recognised as the best automated system for calculating protein-protein interactions.
"In normal cells there are thousands of different protein interactions. Explaining these interactions will help us to describe important processes: how the body works as a whole, and methods of treating certain diseases (such as cancer)," says Dima Kozakov commenting on the study.
Andrey Kazennov, a postgraduate student at MIPT, and Dmitry Padhorny, a postgraduate student at Stony Brook with a Master's degree from MIPT, also contributed to the paper.
###
@phystech_en
Media Contact
Sergey Divakov
[email protected]
@phystech
https://mipt.ru/english/