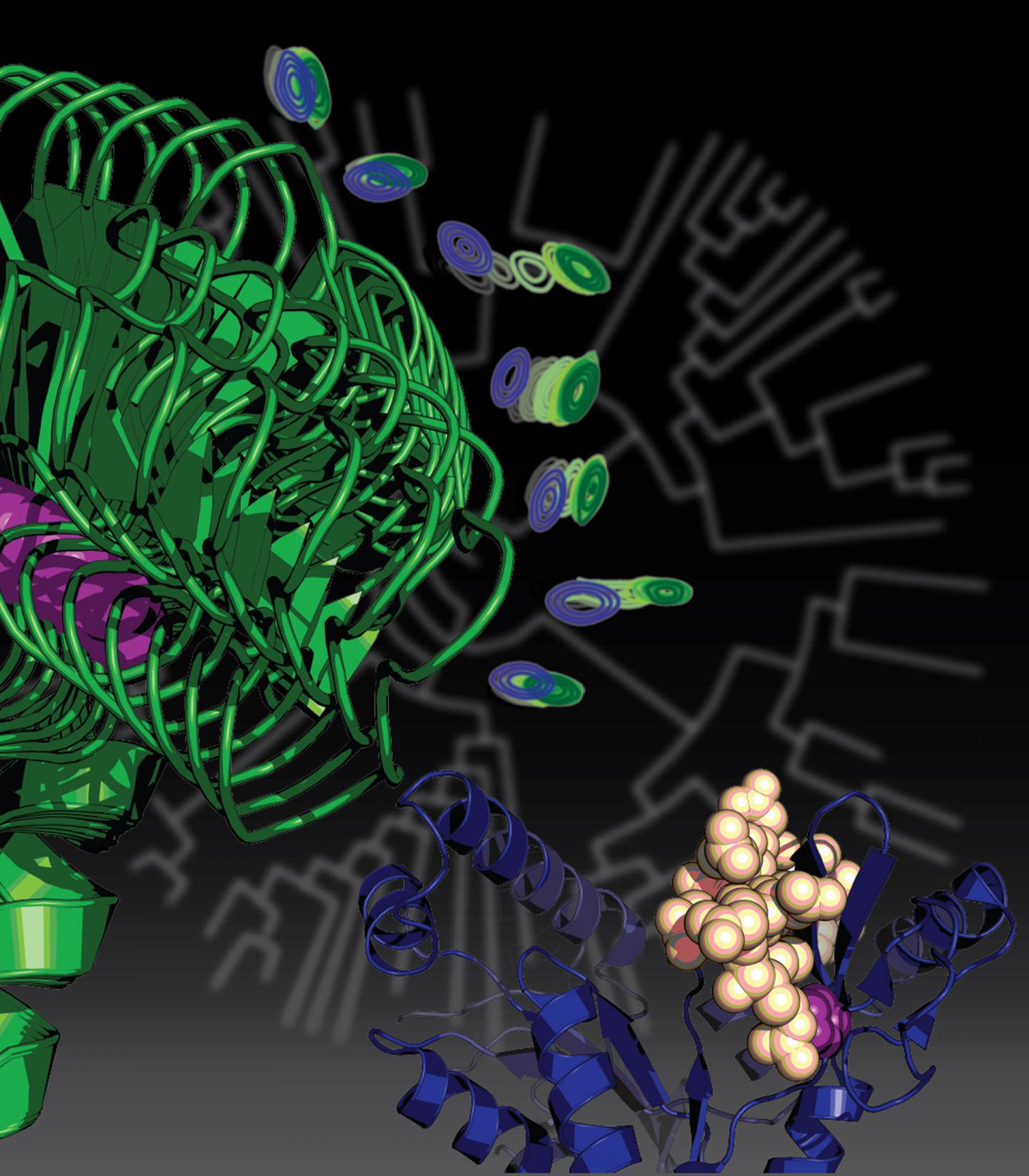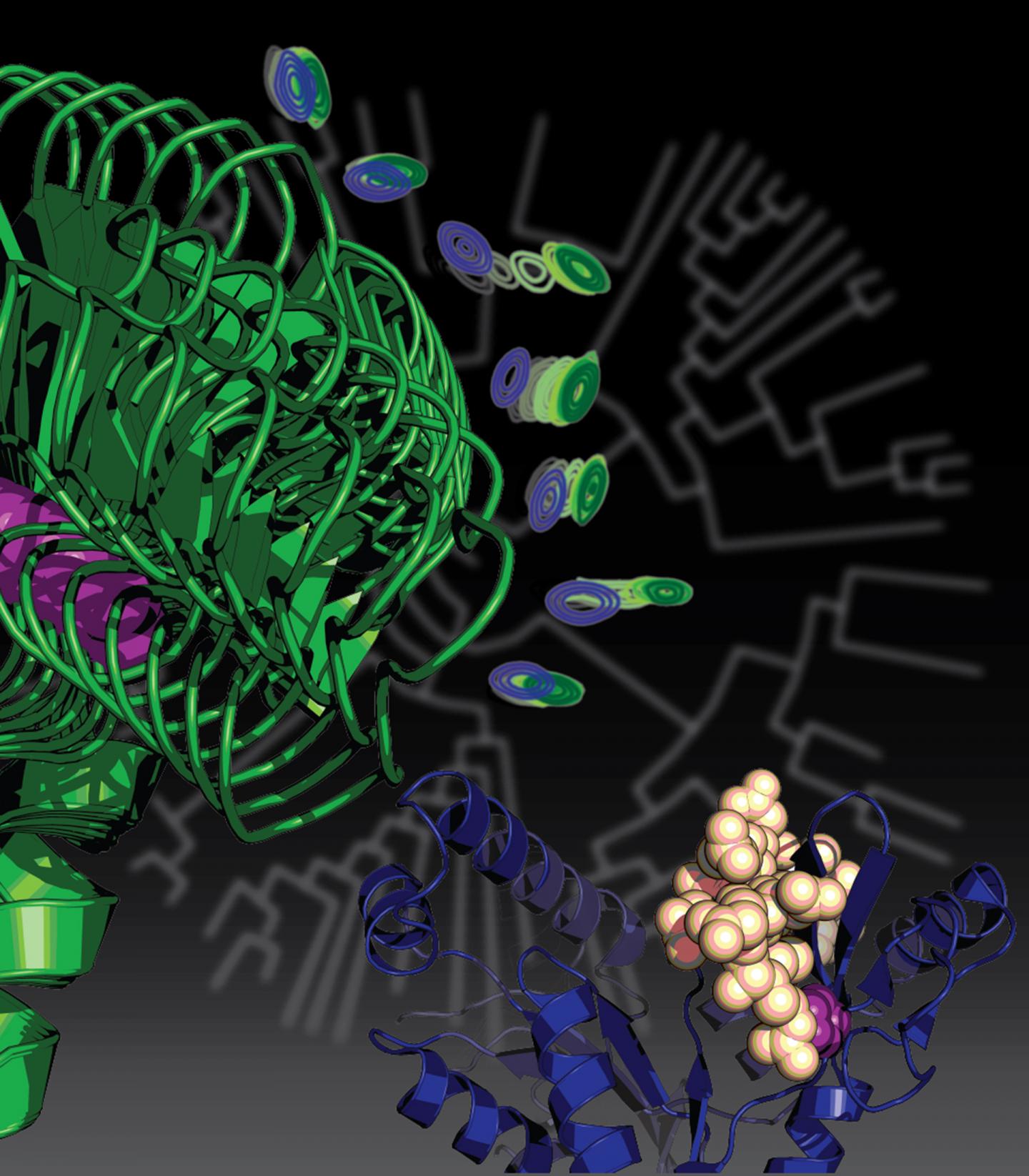
Credit: Courtesy of Kenneth Prehoda
EUGENE, Ore. — Nov. 16, 2016 — Just as a boat can be driven off course by a log in its path, a single, random mutation can send life in a new direction. That scenario, says University of Oregon biochemist Ken Prehoda, illustrates how a random mutation sparked a huge jump in the evolutionary course of a protein important for the evolution of animals.
In January, Prehoda was on a team that found that a random mutation 600 million years ago in a single-celled organism created a new family of proteins that are important for multicellular life. In a new paper, Prehoda and colleagues describe what the mutation did to the original protein, an enzyme known as guanylate kinase.
The paper, now online, will be featured with an illustration on the cover of the Nov. 23 issue of the Journal of the American Chemical Society.
Mutations happen randomly. Most are bad news. Understanding them better, Prehoda said, could potentially point to new treatments for human diseases such as cancer. Occasionally a mutation is good, helping an organism adapt to environmental changes or advancing overall fitness.
Prehoda's lab initially used a molecular technique called ancestral protein reconstruction. The technique allows researchers to move backward in the evolutionary tree to see molecular changes and infer how proteins performed in the past.
For the new study, Prehoda's lab collaborated with researchers at the Medical College of Wisconsin who studied whether the mutation they had discovered had possibly changed the flexibility of the protein. Next, his team turned to computer simulations in the UO's High Performance Computing Research Core Facility to explore how the altered flexibility they isolated, in turn, led to changes in the protein's interactions.
"We found that this mutation that helped our unicellular ancestor to become multicellular, and ended up leading to an entirely new family of proteins that are specific to animals, did so in a very interesting way," said Prehoda, who is the director of the UO's Institute of Molecular Biology. "Amazingly, this one mutation took a protein that was really flexible — an important trait for its old job — and made it much more rigid so it could advance to a new function."
The mutation, which researchers labeled s36P, set off a cascade of events in which guanylate kinase interactions took new routes and evolved into more complex multicellular organisms, Prehoda said. The mutation is still conserved in all animals today, he added.
"A lot of the proteins that do the work in our bodies can be thought of as molecular machines," Prehoda said. "They move in a way that is coordinated with function. Each protein spins in a circle or motors along filaments. Our protein, before the mutation, was an enzyme that had certain flexible movements related to its function. This one mutation fixed the protein's backbone, locking the molecule into a shape that is important for its new function."
Prehoda and colleagues reported their discovery of mutation in a paper that appeared Jan. 7 in the journal eLife.
###
Co-authors on the JACS paper were Dustin S. Whitney and Brian F. Volkman, both of the Medical College of Wisconsin in Milwaukee. They also were among the co-authors on the eLife paper.
Two grants from the National Institutes of Health supported the research.
Source: Ken Prehoda, professor of biochemistry, biophysics & molecular biology, 541-346-5030, [email protected]
Note: The UO is equipped with an on-campus television studio with a point-of-origin Vyvx connection, which provides broadcast-quality video to networks worldwide via fiber optic network. There also is video access to satellite uplink and audio access to an ISDN codec for broadcast-quality radio interviews.
Links:
January news release on the eLife paper: https://www.eurekalert.org/pub_releases/2016-01/uoo-rmp010716.php
Prehoda faculty page: http://chemistry.uoregon.edu/profile/prehoda/
Prehoda lab: http://molbio.uoregon.edu/~prehoda/
Department of Chemistry and Biochemistry: http://chemistry.uoregon.edu/
Institute of Molecular Biology: http://molbio.uoregon.edu/
Media Contact
Jim Barlow
[email protected]
541-346-3481
@UOregonNews
http://uonews.uoregon.edu
############
Story Source: Materials provided by Scienmag