Credit: City University of Hong Kong
Neuronal development is often regulated by the graded distribution of guidance molecules, which can either attract or repel the neuronal migration or neurite projection when presented in a format of concentration gradients, or chemotaxis. However, many details about the process is largely unexplored.
A research team from City University of Hong Kong (CityU) has tackled this problem in a precise and systematic way by developing a new device, and has recently published their findings in a research article on the journal Nature Communications, titled "High-throughput three-dimensional chemotactic assays reveal steepness-dependent complexity in neuronal sensation to molecular gradients".
Chemotaxis refers to the movement of an organism in response to a chemical stimulus. It is well known that the concentration gradients of guidance molecules, such as netrin or semaphorin (Sema) proteins, play critical roles in embryonic neural development. Yet, how exactly the physical profiles of molecular gradients, e.g. the changing rate of concentration profiles (gradient steepness), interplays with neuronal development has long remained an unanswered question. Part of the reason was the lack of 3D devices that can recapitulate important features of brain tissues outside the human body. Previous in vitro chemotactic assays are often 2D, low-throughput (meaning it needs to manually repeat the experiments many times to collect data for different parameters) and lack fine gradient control.
In response, the CityU team develop a new platform for performing the chemotactic experiments. They have developed a hydrogel-based microfluidic platform for high-throughput 3D chemotactic assays, and used it to study neuronal sensitivity to the steepness of molecular gradient, shedding light on neural regeneration mechanism by recognizing subtle variation in the gradient profiles of guidance molecules.
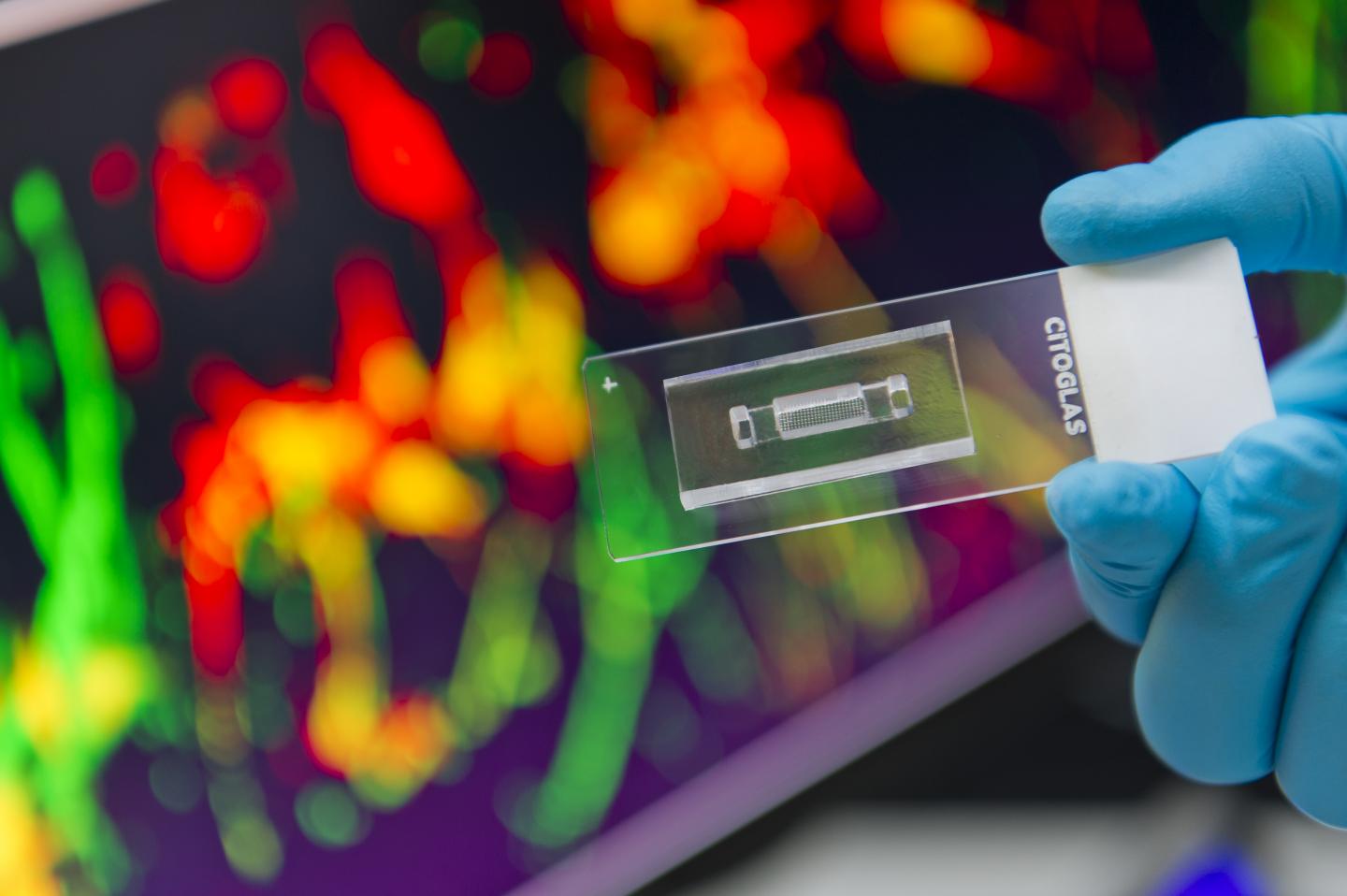
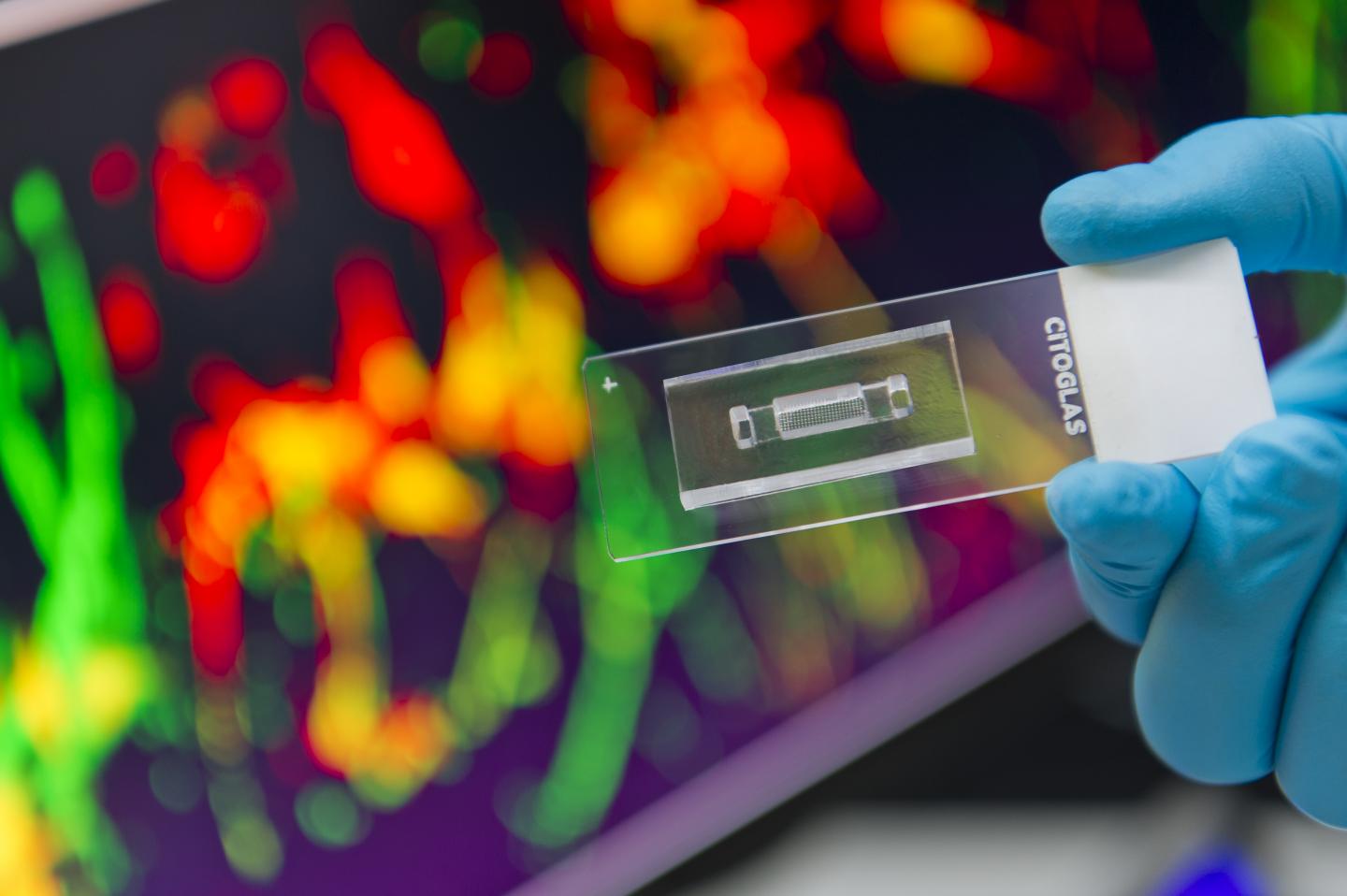
"Our chip measures only 1 by 3 cm2, but houses hundreds of suspended microscale hydrogel cylinders, each containing a distinct gradient profile to allow 3D growth of neuronal cells in an environment closely resembling that inside our brains," says Dr Shi Peng, Associate Professor in the Department of Biomedical Engineering (BME) at CityU, who led the research.
"The major advantage of the setup is the high throughput, meaning a large collection of molecular gradient profiles can be tested in parallel using a single chip to generate a huge amount of data, and the experiment time can be reduced from months to 48 hours," he explains.
Using the new platform and rigorous statistical analysis, the team has revealed dramatic diversity and complexity in the chemotactic regulation of neuronal development by various guidance molecules. In particular, for Sema3A, the team has found that two signaling pathways, namely STK11 and GSK3, are differentially involved in steepness-dependent chemotactic regulation of coordinated neurite repellence and neuronal migration.
Based on these findings, the team further demonstrated that the guidance molecule, Sema3A, is only beneficial to promote cortex regeneration if it is presented in the right gradient form in an injured rat brain, which was reported in another article on the journal Biomaterials earlier this year.
"In case of brain injury, the nervous system does not regenerate easily, so proper use of guidance molecules would help the brain to recover. In this regard, our research provides insights to the development of novel therapeutic strategies," Dr Shi concluded.
###
The first co-authors of the paper are Xu Zhen, Fang Peilin and Xu Bingzhe from BME. The other authors include Dr Fan Jun, Lu Yufeng and Xiong Jinghui from Department of Material Science and Engineering, and Dr Wang Xin and Gao Feng from Department of Biomedical Science at CityU.
Media Contact
P.K. Lee
[email protected]
852-344-28925
https://www.cityu.edu.hk/
Related Journal Article
http://dx.doi.org/10.1038/s41467-018-07186-x