Credit: University of Lincoln
Scientists will connect two of the most important scientific theories of all time – the law of universal gravitation and the theory of evolution – by unravelling the genetic code of the apple which inspired Isaac Newton's greatest discovery 350 years ago this year.
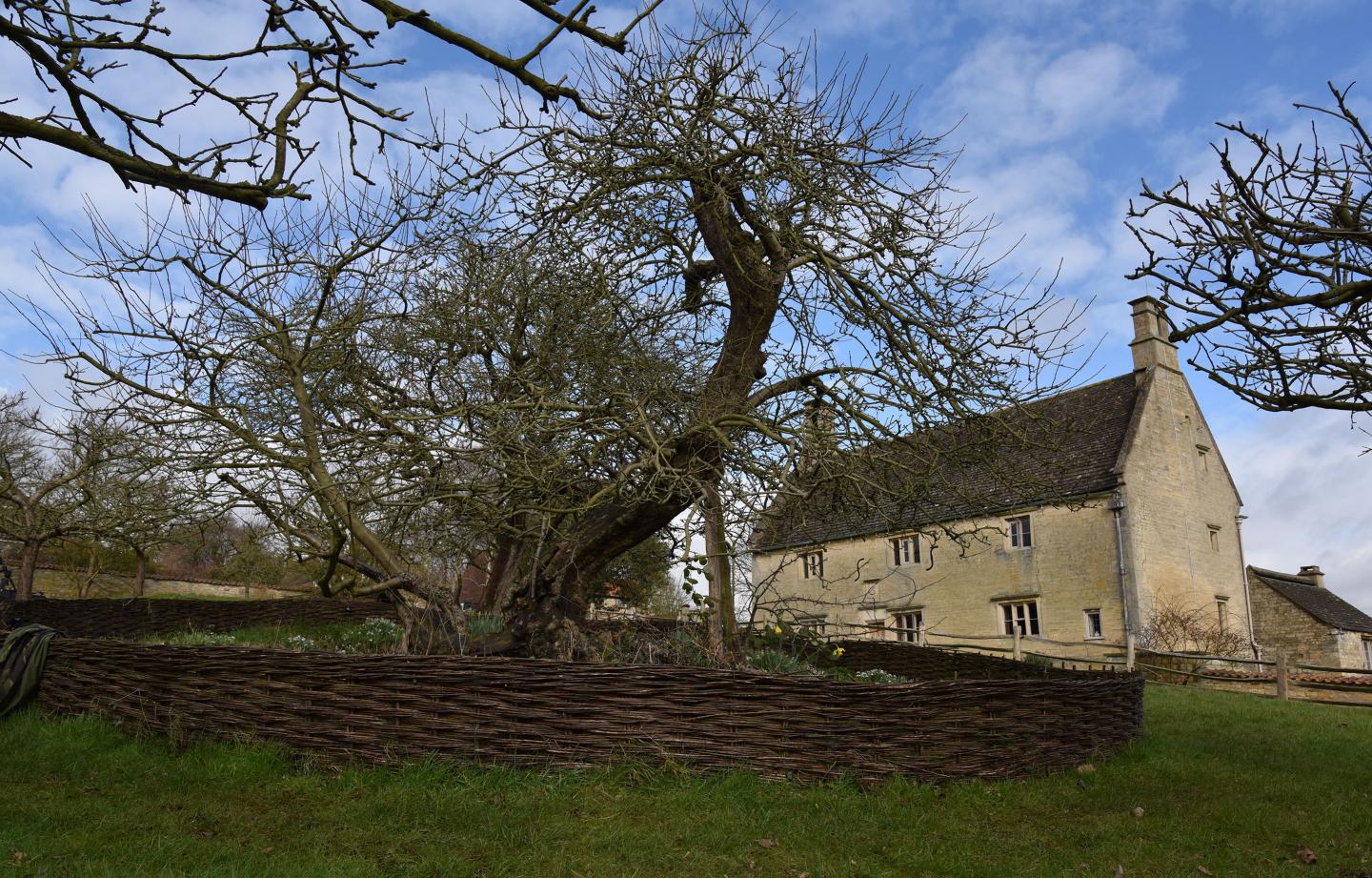
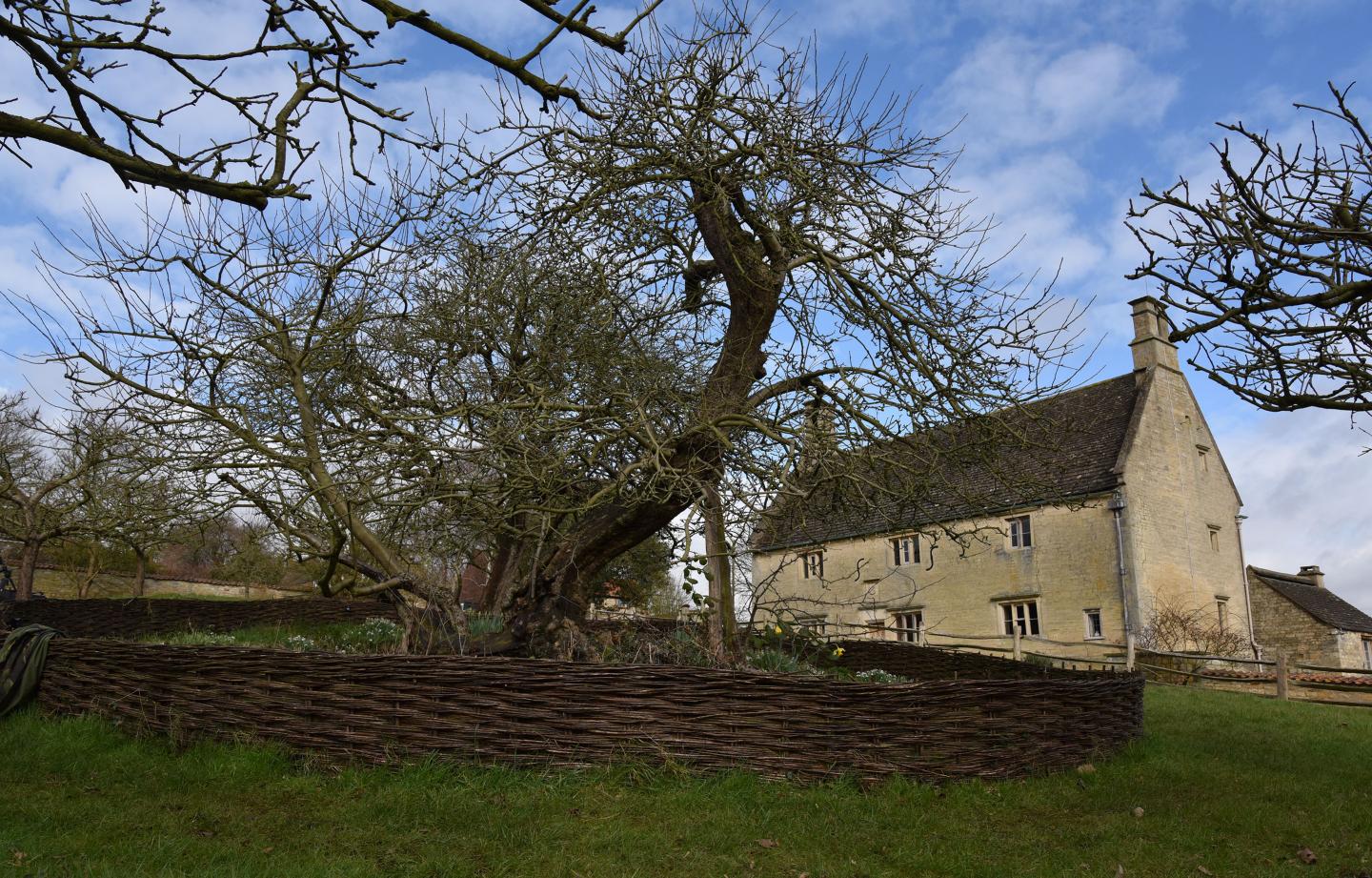
For the first time, researchers are conducting whole genome sequencing of the ancient tree in the garden of Woolsthorpe Manor, Lincolnshire, where Newton is said to have to have seen an apple fall to the ground, prompting him to speculate on the nature of gravity in late 1666.
This period of Newton's life, when his momentous ideas about the laws of motion and gravity were developed, has become known as his Annus mirabilis (year of wonders) and is considered a pivotal moment in the Age of Enlightenment.
The whole genome sequencing of Newton's apple is being carried out by scientists from the University of Lincoln, UK, and the Earlham Institute (EI) (formerly The Genome Analysis Centre), UK. Their goal is to gain a deeper understanding of how technological and societal changes triggered by the scientific revolution Newton's work made possible have affected the evolution of apples over the past 350 years.
Professor Matthew Goddard, an evolutionary biologist in the School of Life Sciences at the University of Lincoln, is leading the research. He said: "Sequencing the genome of Newton's apple connects two of the most influential thinkers in the history of science: Isaac Newton and Charles Darwin. It creates an intersection between these two fundamental theories of gravity and evolution."
The genome of an organism contains the complete set of its DNA. Darwin's theory of evolution, set out in his defining work On the Origin of Species (1859), established natural selection as the principal force in driving rapid genetic variation within a species. But chance errors in DNA replication can also lead to genetic changes occurring over many generations. This process, called neutral evolution, is another powerful force in shaping changes at a molecular level within an organism.
Whole genome sequencing enables even the smallest genetic variations to be analysed.
Professor Goddard, a member of the University's Lincoln Institute for Agri-food Technology (LIAT), explained: "The genome of Newton's apple pre-dates the agricultural revolution. Now we have sequenced it, we can compare it to the genomes of modern apples and see how much the genomes have changed over the centuries. We can then compare that to changes in other organisms to gain a relative understanding of how the pace of human progress since Newton's time has affected the pace of biological change in this particular plant species."
The project is funded by EI and the University of Lincoln. The genome has been sequenced from a graft of the tree donated by The National Trust, owners of Woolsthorpe Manor, to the University of Lincoln earlier this year. Once nurtured, the tree will be planted at the University's Isaac Newton Building, home to staff and students in engineering, computer science, mathematics and physics.
The whole genome sequencing of crop genomes is still in its infancy due to the complexity of plants when compared to human DNA. In order to complete this project, state-of-the-art DNA sequencers available at the Earlham Institute were required to generate the apple tree's data. Sequencing DNA involves 'shredding' DNA into small, overlapping fragments, determining the order of the molecular base pairs that make up the DNA in those fragments and then comparing them to a reference sequence; in this case a modern apple tree cultivar.
Dr Daniel Swan, EI's Head of Platforms and Pipelines, added: "The differences between the modern crop tree, and that of Newton's apples tells a story of apple crop improvement, and allows us to pinpoint those genetic changes that drive the evolution of a crop over time, not only from natural selection, but also human induced selection from breeding and grafting programmes. The apples, however, still fall to the floor under the same gravitational constraints elucidated 350 years ago."
###
Media Contact
Elizabeth Allen
[email protected]
01-522-837-650
@unilincoln
http://www.lincoln.ac.uk/home/
############
Story Source: Materials provided by Scienmag