Credit: Wilfried Guiblet, Penn State
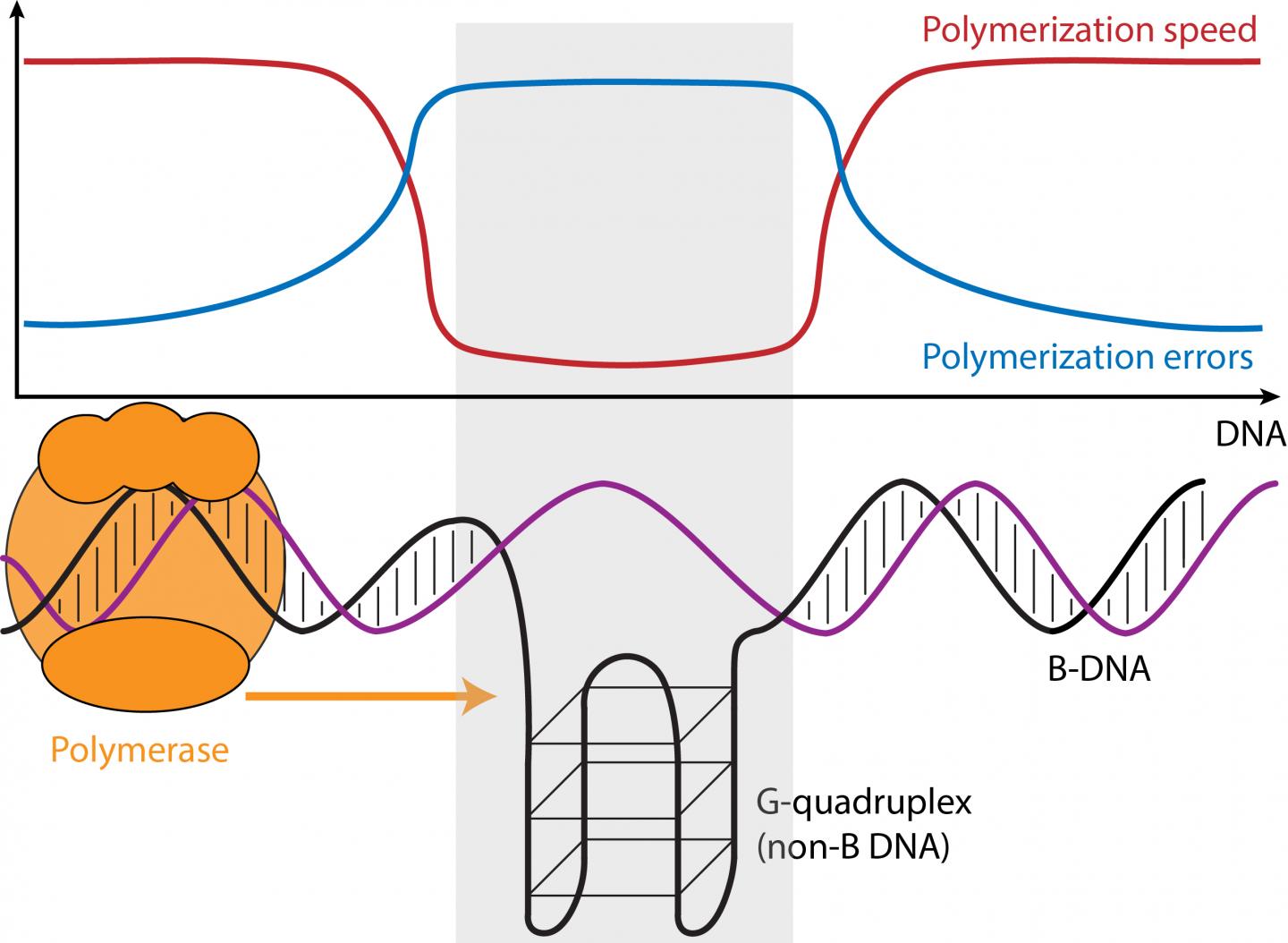
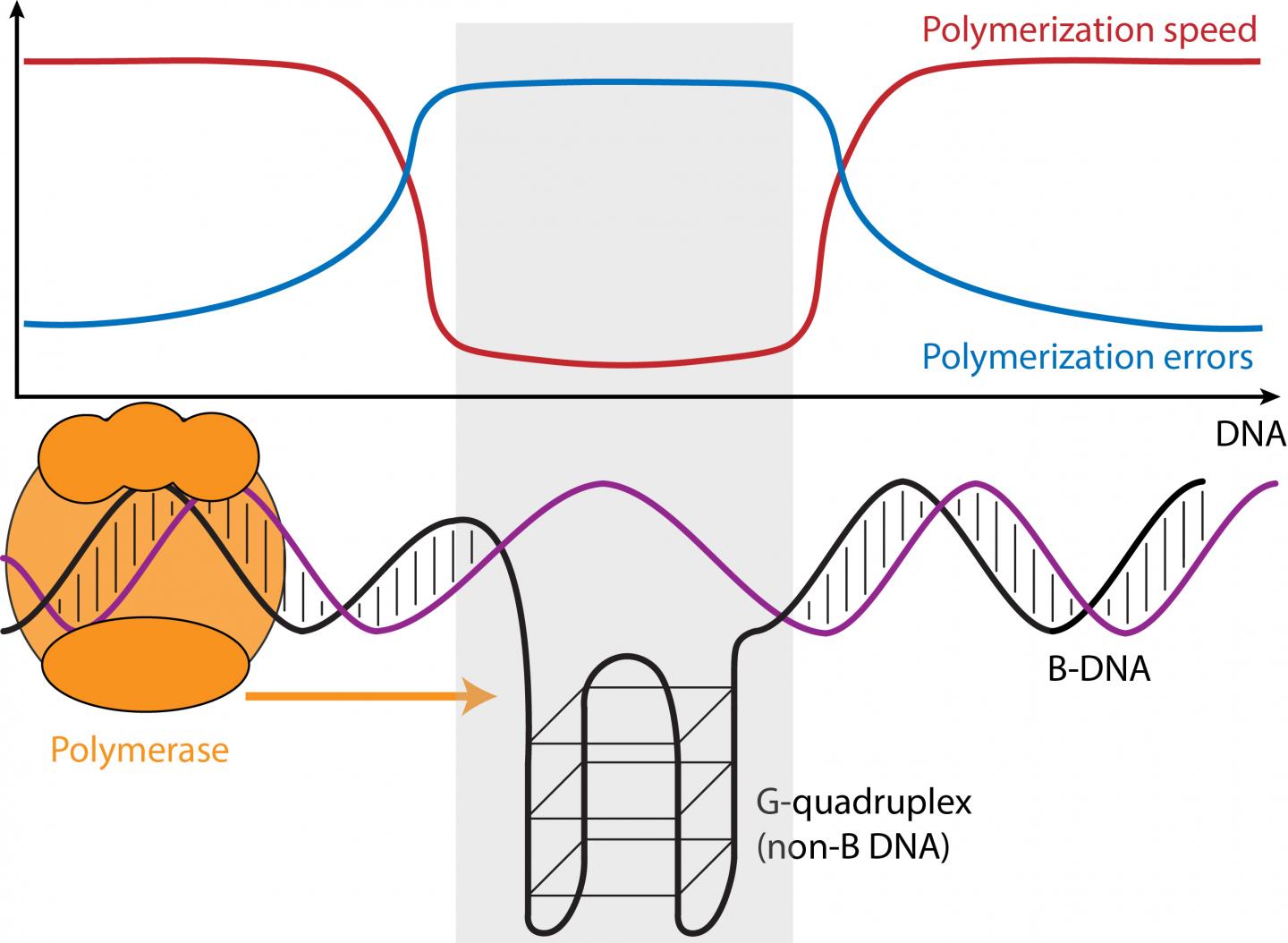
The speed and error rate of DNA synthesis is influenced by the three-dimensional structure of the DNA. Using "third-generation" genome-wide DNA sequencing data, a team of researchers from Penn State and the Czech Academy of Sciences showed that sequences with the potential to form unusual DNA conformations, which are frequently associated with cancer and neurological diseases, can in fact slow down or speed up the DNA synthesis process and cause more or fewer sequencing errors. An article describing the study appears online in the journal Genome Research.
"We've been interested for a long time in trying to understand the factors that affect variation in mutation rates across the genome," said Kateryna Makova, Pentz Professor of Biology at Penn State and one of the leaders of the research team. "Sequences that can form non-B DNA, which form structures other than the common right-handed double-helix with ten bases per turn (B-DNA), make up about 13 percent of the human genome and play many important roles in cellular functions, including gene regulation and the protection of telomeres–the sequences that cap and stabilize the ends of chromosomes. Because these regions are also associated with many human diseases we were interested to see if they had any impact on the speed of the DNA synthesis reaction–also called "polymerization"–and on its error rates."
Non-B DNA includes sequences with runs of the "G" nucleotide, guanine, which can form G-quadruplex structures; "A"-rich regions, which can cause helix bending; and short tandem repeats–regions with the same one-to-six nucleotide motifs repeated over and over again (e.g. GATA GATA GATA …)–that can form slipped strands and hairpins. Using Single-Molecule-Real-Time sequencing, or SMRT, which tracks the time between the incorporation of each successive nucleotide (the A, T, C, or G building blocks of DNA), during sequencing, the researchers compared regions of non-B DNA to B-DNA.
"There are hundreds of thousands of sequence motifs that are predicted to form non-B DNA across the genome," said Wilfried Guiblet, a graduate student in the bioinformatics and genomics program at Penn State and co-first author of the paper. "We used data from the SMRT sequencer from Pacific Biosciences to compare the nucleotide incorporation times along non-B DNA regions with those along regions of more common B-DNA." The comparison revealed that some forms of non-B DNA–G-quadruplexes, for example–caused the polymerase enzyme to slow down by as much as 70 percent, while other non-B DNA caused the enzyme to speed up.
"In order to analyze the data, we developed a novel Functional Data Analysis (FDA) statistical tool," said Francesca Chiaromonte, professor of statistics at Penn State and another leader of the research team. "This tool contrasts the nucleotide incorporation times in non-B and B-DNA regions treating them as curves, or mathematical functions." Marzia Cremona, Bruce Lindsay Visiting Assistant Professor of Statistics at Penn State and another co-first author of the paper, was instrumental in developing the new tool and performing the statistical analysis. A software package implementing the testing procedure is now publicly available.
In addition to changes in nucleotide incorporation times, the researchers also noted that the error rates of the sequencer increased in some types non-B DNA regions, for instance in G-quadruplex motifs. In these regions, the increased error rates lined up with increased DNA sequence variation among humans (using data from the "1000 Genomes Project"), as well as increased divergence between human and orangutan.
"To perform sequencing, SMRT uses an enzyme called polymerase, similar to the cell using polymerases to replicate DNA," said Makova. "It seems likely that the same phenomenon that is causing the increased error rate in the sequencer is also causing the increase we see in human variation and divergence from the orangutan. Understanding how the structure of non-B DNA impacts mutation rates is extremely interesting from the standpoint of genome evolution, and also because these regions have been implicated in human diseases."
###
In addition to Makova, Guiblet, Cremona, and Chiaromonte, the research team includes Monika Cechova, Robert S. Harris, and Kristin Eckert at Penn State; and Iva Kejnovska and Eduard Kejnovsky at the Academy of the Sciences of the Czech Republic. The research was funded by the Eberly College of Sciences, the Huck Institutes of the Life Sciences, and the Institute for CyberScience, at Penn State; by grants from the Pennsylvania Department of Health; and by the Czech Science Foundation.
Media Contact
Sam Sholtis
[email protected]
814-865-1390
@penn_state
http://live.psu.edu
Original Source
http://science.psu.edu/news-and-events/2018-news/Makova11-2018 http://dx.doi.org/10.1101/gr.241257.118