Computer programs have defeated humans in Jeopardy!, chess and Go. Now a program developed at Case Western Reserve University has outperformed physicians on a more serious matter.
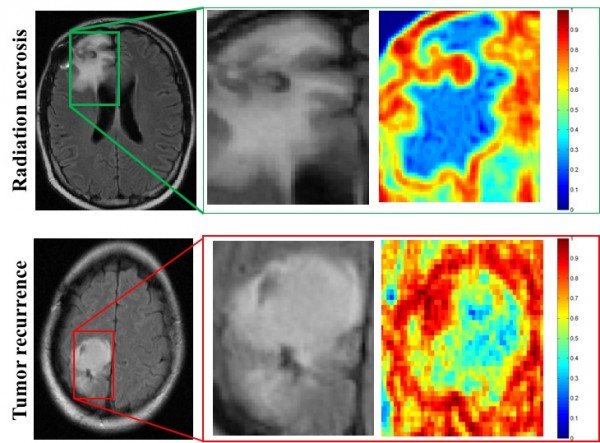
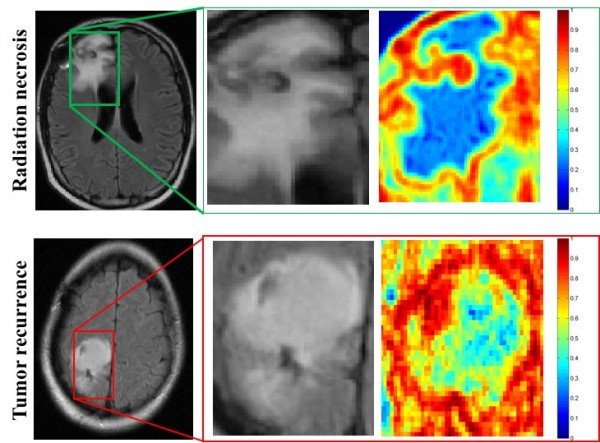
The program was nearly twice as accurate as two neuroradiologists in determining whether abnormal tissue seen on magnetic resonance images (MRI) were dead brain cells caused by radiation, called radiation necrosis, or if brain cancer had returned.
The direct comparison is part of a feasibility study published in the American Journal of Neuroradiology today.
“One of the biggest challenges with the evaluation of brain tumor treatment is distinguishing between the confounding effects of radiation and cancer recurrence,” said Pallavi Tiwari, assistant professor of biomedical engineering at Case Western Reserve and leader of the study. “On an MRI, they look very similar.”
But treatments for radiation necrosis and cancer recurrence are far different. Quick identification can help speed prognosis, therapy and improve patient outcomes, the researchers say.
With further confirmation of its accuracy, radiologists using their expertise and the program may eliminate unnecessary and costly biopsies Tiwari said. Brain biopsies are currently the only definitive test but are highly invasive and risky, causing considerable morbidity and mortality.
To develop the program, the researchers employed machine learning algorithms in conjunction with radiomics, the term used for features extracted from images using computer algorithms. The engineers, scientists and physicians trained the computer to identify radiomic features that discriminate between brain cancer and radiation necrosis, using routine follow-up MRI scans from 43 patients. The images were all from University Hospitals Case Medical Center.
The team then developed algorithms to find the most discriminating radiomic features, in this case, textures that can’t be seen by simply eyeballing the images.
“What the algorithms see that the radiologists don’t are the subtle differences in quantitative measurements of tumor heterogeneity and breakdown in microarchitecture on MRI, which are higher for tumor recurrence,” said Tiwari, who was appointed to the Department of Biomedical Engineering by the Case Western Reserve School of Medicine.
More specifically, while the physicians use the intensity of pixels on MRI scans as a guide, the computer looks at the edges of each pixel, explained Anant Madabhushi, F. Alex Nason professor II of biomedical engineering at Case Western Reserve, and study co-author.
“If the edges all point to the same direction, the architecture is preserved,” said Madabhushi, who also directs the Center of Computational Imaging and Personalized Diagnostics at CWRU. “If they point in different directions, the architecture is disrupted — the entropy, or disorder, and heterogeneity are higher. ”
In the direct comparison, two physicians and the computer program analyzed MRI scans from 15 patients from University of Texas Southwest Medical Center. One neuroradiologist diagnosed seven patients correctly, and the second physician correctly diagnosed eight patients. The computer program was correct on 12 of the 15.
Tiwari and Madabhushi don’t expect the computer program would be used alone, but as a decision support to assist neuroradiologists in improving their confidence in identifying a suspicious lesion as radiation necrosis or cancer recurrence.
Next, the researchers are seeking to validate and the algorithms’ accuracy using a much larger collection of images from across different sites.
Story Source:
The above post is reprinted from materials provided by Case Western Reserve University.
Journal Reference:
P. Tiwari, P. Prasanna, L. Wolansky, M. Pinho, M. Cohen, A.P. Nayate, A. Gupta, G. Singh, K. Hattanpaa, A. Sloan, L. Rogers, and A. Madabhushi. Computer-Extracted Texture Features to Distinguish Cerebral Radionecrosis from Recurrent Brain Tumors on Multiparametric MRI: A Feasibility Study. American Journal of Neuroradiology, September 2016 DOI: 10.3174/ajnr.A4931
The post Computer program beats physicians at brain cancer diagnoses appeared first on Scienmag.