Single-molecule approach to RNA transcription brings new details to light
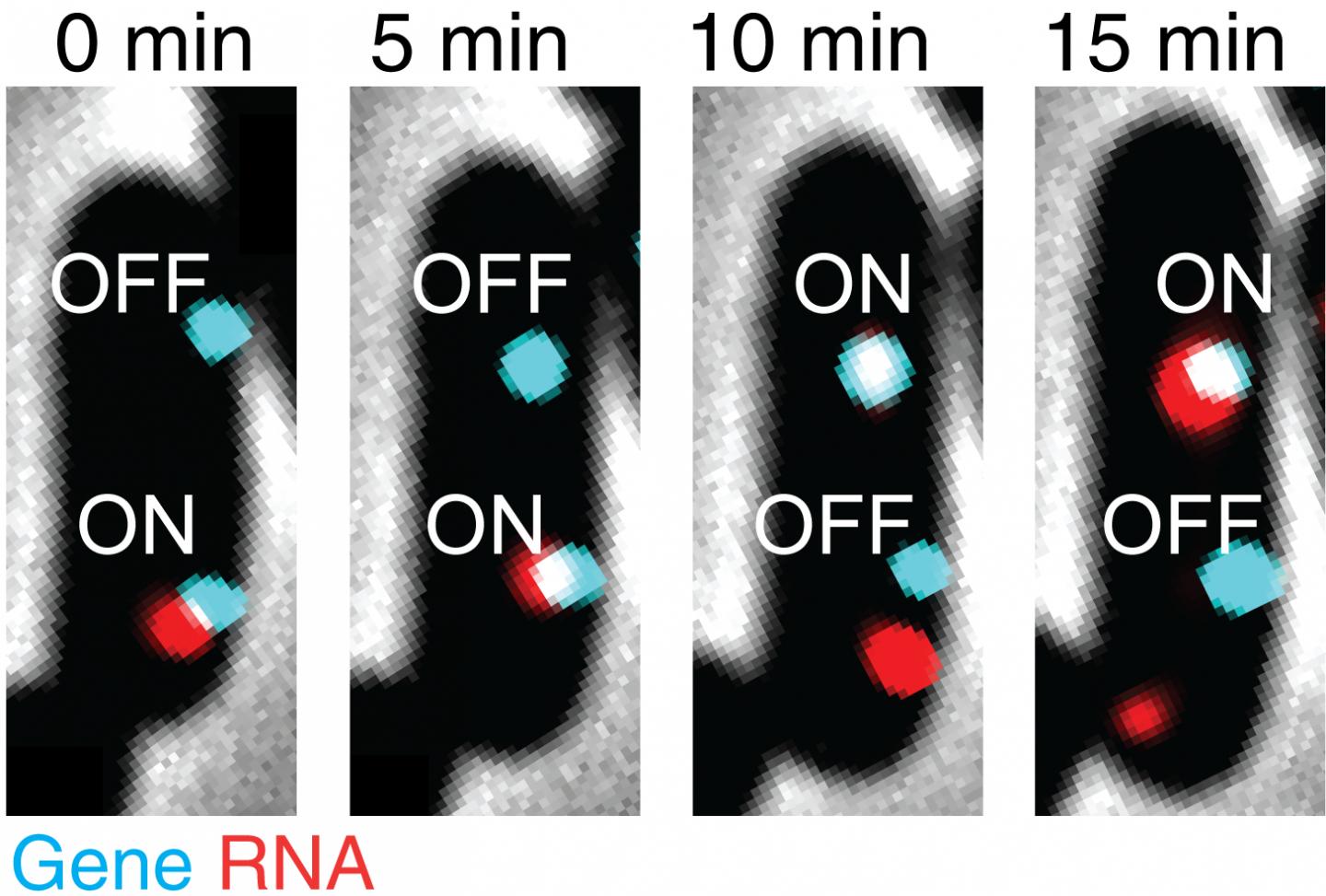
Credit: Mengyu Wang, Jing Zhang, Heng Xu, and Ido Golding
Scientists studying genetic transcription are gaining new insights into a process that is fundamental to all life. Transcription is the first step in gene expression, the process taking place within all living cells by which the DNA sequence of a gene is copied into RNA, which in turn (most generally speaking) serves as the template for assembling protein molecules, the basic building blocks of life.
Much of what scientists have uncovered about transcription over the past five decades is based on bulk investigative techniques employing large numbers of living cells. Today, advanced imaging techniques allow scientists to probe the inner workings of transcription at the scale of individual genes, and a new more detailed picture of this vital process is emerging.
Just this week, two new in vivo single-molecule studies of transcription in E. coli were published by scientists at the University of Illinois at Urbana-Champaign, one by Professor Ido Golding and colleagues, unveiling unexpected and up-to-now hidden drivers of cellular individuality; the other by Professor Sangjin Kim and colleagues, demonstrating for the first time that transcription dynamics are affected by molecular-scale long-distance communication between RNA polymerase (RNAP) molecules while they are “reading” a gene sequence one base at a time and assembling the complementary RNA strand.
Taken together, these two studies elucidate new details of the physical processes of gene expression at the level of individual cells and initiate an exciting new direction of inquiry for member scientists of the Center for the Physics of Living Cells (CPLC), a National Science Foundation Physics Frontiers Center in the Department of Physics at the University of Illinois at Urbana-Champaign.
Kim notes, “My dream project is to probe the physical mechanism of this emergent phenomenon–the coupling between DNA supercoiling and RNA polymerase motion. The CPLC is a fantastic place to pursue this and to collaborate with other theoretical and experimental scientists at the intersection of microbiology, biochemistry, and biophysics. I have already connected with wonderful new colleagues who are interested in collaborating on this, including Ido Golding, Nigel Goldenfeld, and Yann Chemla.”
Golding and Kim only recently joined the faculty at Illinois Physics, and the two unrelated but complementary experiments were performed at the scientist’s respective prior institutions. Golding, who had been a faculty member at Illinois Physics from 2007 to 2009, returned to Illinois Physics in July 2019 from Baylor College of Medicine in Houston, where he held an appointment as a professor of biochemistry and molecular biology. Kim joined the faculty at Illinois Physics in January 2019, following a postdoctoral appointment at Yale University, where she worked in the research group of noted microbiologist Professor Christine Jacobs-Wagner.
Drivers of bacterial individuality
Golding, along with colleagues at Baylor College of Medicine and Shanghai Jiao Tong University in China, have uncovered specific drivers of bacterial individuality, ultimately bringing the process of transcription into greater focus. The researchers used single-cell measurements as well as computational modeling to characterize the phases of gene transcription in terms of their dynamics.
The team’s findings were published online in a Nature Microbiology letter on September 16, 2019.
The team observed that a weakly expressed gene exhibits a temporary pulse of transcription activity around the time of gene replication. In other words, transcription responds, directly or indirectly, to the event of gene replication.
“It is too soon to determine definitively what the underlying mechanism is for this coupling of DNA replication and RNA transcription,” notes Golding. “But it is clear that this phenomenon is a driver of individuality among cells in a colony, because individual cells in the growing population are unsynchronized in their cell-cycle phase, therefore each will replicate the gene at a different time.”
Golding adds, “While we still do not know the reason for this coupling of transcription and gene replication, there are many plausible hypotheses, and theorists have been speculating on this for many years. It’s been suggested, for example, that the newly synthesized DNA–the gene–is more susceptible to binding by the cellular machines that drive transcription (RNAP and others).”
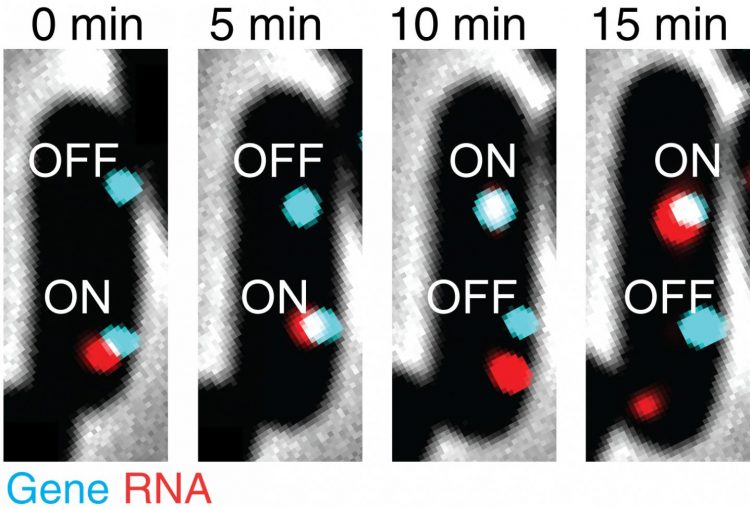
Golding’s postdoctoral researcher Mengyu Wang, who is co-lead author on this study, followed Golding to Illinois and the CPLC. Wang remembers his initial surprise when repeated experimental runs yielded another transcription correlation. The researchers found that when there are two or more copies of the same gene within the same cell, they can sometimes affect each other, switching transcription on or off in unison.
Wang comments, “Scientists had assumed there is no correlation between different gene copies. We repeated our experiment several times and, within each run, there were a few samples, so we are confident in our result. There exists a correlation between different copies of the same gene, and this correlation depends on the growth conditions. Further investigation is necessary to understand the underlying mechanism at work here.”
Golding notes that there is still a lot of ground to cover in our investigations of RNA transcription, to bring the process into greater focus.
“Theoretical models that both interpret and predict the results of
bulk studies represent the intracellular process of transcription as stochastic–it’s inherently random and unpredictable at the level of the cell. These models are very useful to our understanding of what’s happening both at the level of individual cells and within colonies of cells. But such course-grained pictures run the risk of labelling everything that’s unknown as unknowable and random,” remarks Golding.
New research initiatives at the Center for the Physics of Living Cells promise to help uncover the specifics of the most fundamental biological processes involved in heredity and the uniqueness of individuals.
“By examining transcription as it takes place at an individual copy of a single gene in the cell, rather than making deductions from the overall number of RNA molecules present in the cell, we were able to observe new details that current theoretical models of the process don’t take into account,” Golding concludes.
At Illinois, Golding is a member of the CPLC and is an affiliate of the Carl R. Woese Institute for Genomic Biology and of the Department of Microbiology.
This work was supported by the National Institutes of Health, the National Science Foundation, and by grants from private foundations and Chinese national funding agencies. The conclusions presented are those of the researchers and not necessarily those of the funding agencies.
Long-range “communication” among RNA polymerase molecules on a single gene
Kim, along with colleagues at the interdisciplinary Microbial Sciences Institute at Yale University, performed in vitro and in vivo experiments on genes having multiple RNAPs simultaneously transcribing the gene, each at different points of progress along the gene. The team observed long-range interactions that accelerated or impaired neighboring RNAPs’ transcription speed, depending on whether the gene was turned on or off in response to environmental conditions.
These findings were published online on September 19, 2019 in the journal Cell.
The team observed that, surprisingly, RNAPs can communicate with each other from distances as far as two thousand bases apart on the DNA. And when there are multiple RNAPs, they translocate faster than a single RNAP.
Kim explains the mechanism underlying this communication.
“This communication is coming from a property of DNA called supercoiling, which dynamically changes during both replication and transcription,” she describes.
“DNA is a double helix that supercoils–like a rope that can be twisted–in response to a portion of the double strand being opened so that it can be read,” Kim continues. “The twisting is a natural consequence of the function. In transcription, as RNAP translocates along the DNA, DNA becomes twisted, but our study shows that this in turn becomes a mechanism allowing for the long-distance communication between RNAPs.”
The team further found that when the promoter–the portion of a gene that acts like a switch to turn transcription on and off, depending on environmental conditions and cellular needs–was switched off, RNAPs slow down and some become disconnected from the gene.
According to Kim, further study of the collective behavior of RNAPs could shed new light on a variety of molecular processes happening on the DNA, including the evolution of gene mutations.
“It is known that when RNAP stops, it can introduce mutations on the genetic code. This has possible implications for the rise of antibiotic resistance in bacteria. In future work at the CPLC, I want to test this by modulating DNA supercoiling and looking deeper at the mechanics. This line of inquiry provides a great opportunity for collaboration between theory and experiment.”
###
Media Contact
Ido Golding
[email protected]
Original Source
https:/
Related Journal Article
http://dx.