Credit: Credit: Purdue Agricultural Communications/Tom Campbell
WEST LAFAYETTE, Ind. – Doctors may soon be able to detect and monitor a patient's cancer with a simple blood test, reducing or eliminating the need for more invasive procedures, according to Purdue University research.
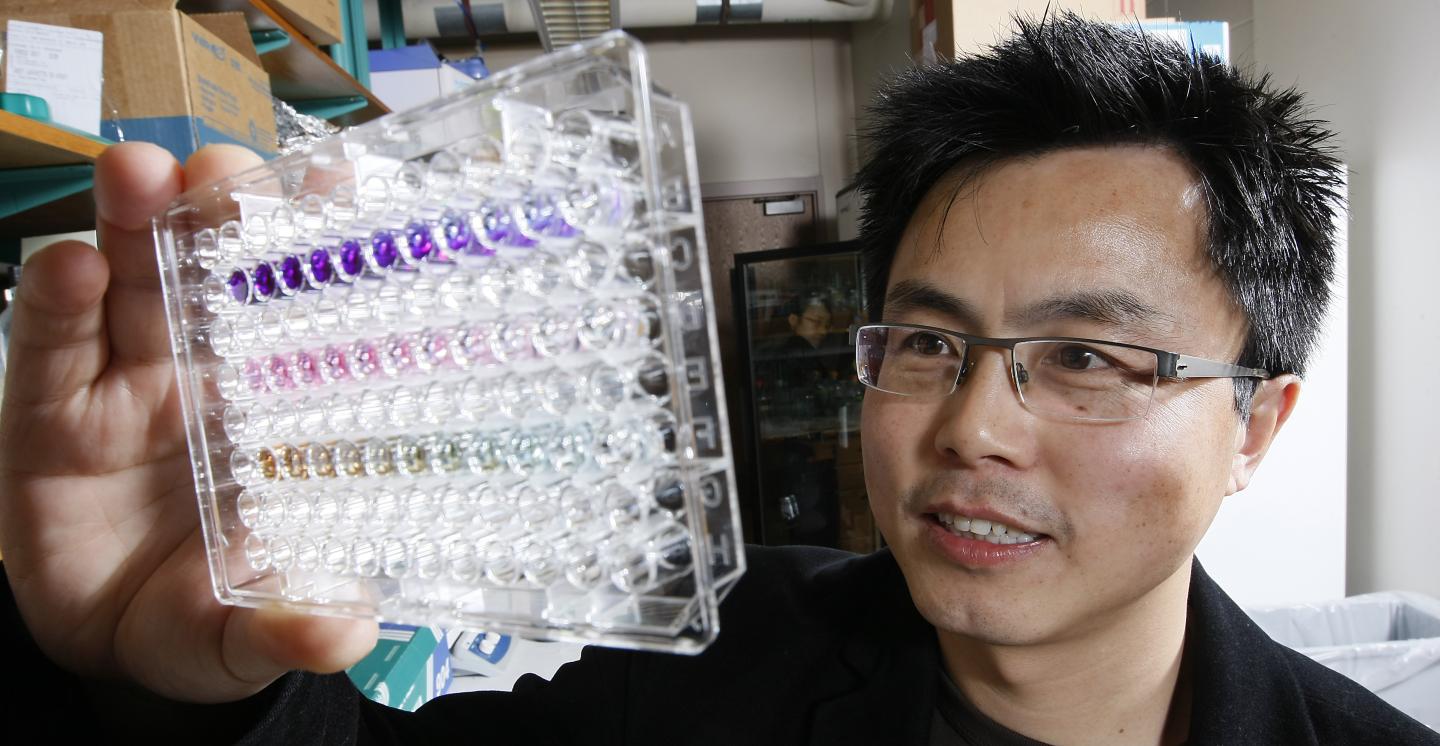
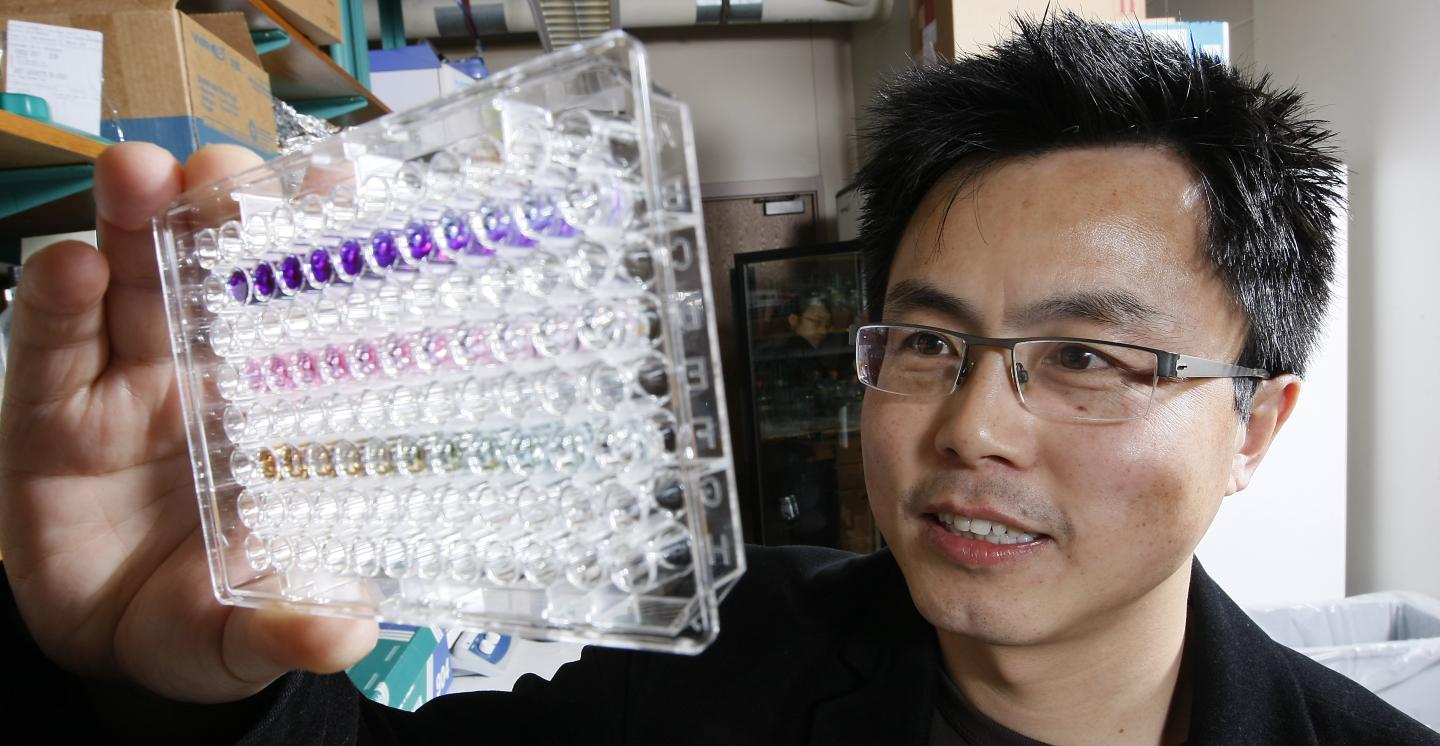
W. Andy Tao, a professor of biochemistry and member of the Purdue University Center for Cancer Research, and colleagues identified a series of proteins in blood plasma that, when elevated, signify that the patient has cancer. Their findings were published in the early edition of the Proceedings of the National Academy of Sciences.
Tao's work was done with samples from breast cancer patients, but it is possible the method could work for any type of cancer and other types of diseases. The work relies on analysis of microvesicles and exosomes in blood plasma.
Protein phosphorylation, the addition of a phosphate group to a protein can lead to cancer cell formation. So phosphorylated proteins, known as phosphoproteins, have been seen as prime candidates for cancer biomarkers. Until now, however, scientists weren't sure identification of phosphoproteins in blood was possible because the liver releases phosphatase into the bloodstream, which dephosphorylates proteins.
"There are so many types of cancer, even multiple forms for different types of cancer, that finding biomarkers has been discouraging," Tao said. "This is definitely a breakthrough, showing the feasibility of using phosphoproteins in blood for detecting and monitoring diseases."
Tao and his colleagues found nearly 2,400 phosphoproteins in a blood sample and identified 144 that were significantly elevated in cancer patients. The study compared 1-milliliter blood samples from 30 breast cancer patients with six healthy controls.
The researchers used centrifuges to separate plasma from red blood cells, and high-speed and ultra-high-speed centrifuges to further separate microvesicles and exosomes. Those particles, which are released from cells and enter the bloodstream, may play a role in intercellular communication and are thought to be involved in metastasis, spreading cancer from one place to another in the body. They also encapsulate phosphoproteins, which Tao's team identified using mass spectrometry.
"Extracellular vesicles, which include exosomes and microvesicles, are membrane-encapsulated. They are stable, which is important," Tao said. "The samples we used were 5 years old, and we were still able to identify phosphoproteins, suggesting this is a viable method for identifying disease biomarkers."
A simple blood test for cancer would be far less invasive than scopes or biopsies that remove tissue. A doctor could also regularly test a cancer patient's blood to understand the effectiveness of treatment and monitor patients after treatment to see if the cancer is returning.
"There is currently almost no way to monitor patients after treatment," Tao said. "Doctors have to wait until cancer comes back."
Timothy Ratliff, director of the Purdue University Center for Cancer Research, said the findings are promising for early detection of cancer.
"The vesicles and exosomes are present and released by all cancers, so it could be that there are general patterns for cancer tissues, but it's more likely that Andy will develop patterns associated with different cancers. It's really exciting," Ratliff said. "Early detection in cancer is key and has been shown to clearly reduce the death rate associated with the disease."
Tao plans to analyze increased levels of phosphoproteins in various types of cancer to determine whether there are patterns that would signify the type of cancer a patient has. His company, Tymora Analytical, is also developing technology that would allow doctors to insert blood samples onto a cartridge and analyze phosphoproteins present, eliminating the need for ultra-high-speed centrifuges that aren't practical in clinical settings.
###
The National Institutes of Health, the National Science Foundation and the Purdue University Center for Cancer Research supported Tao's research.
Contact: Shari Finnell, 317-201-2345, [email protected]
Media Contact
Shari L Finnell
[email protected]
317-201-2345
@PurdueUnivNews
http://www.purdue.edu/
############
Story Source: Materials provided by Scienmag