New insights into ribosome recycling with enzyme ABCE1
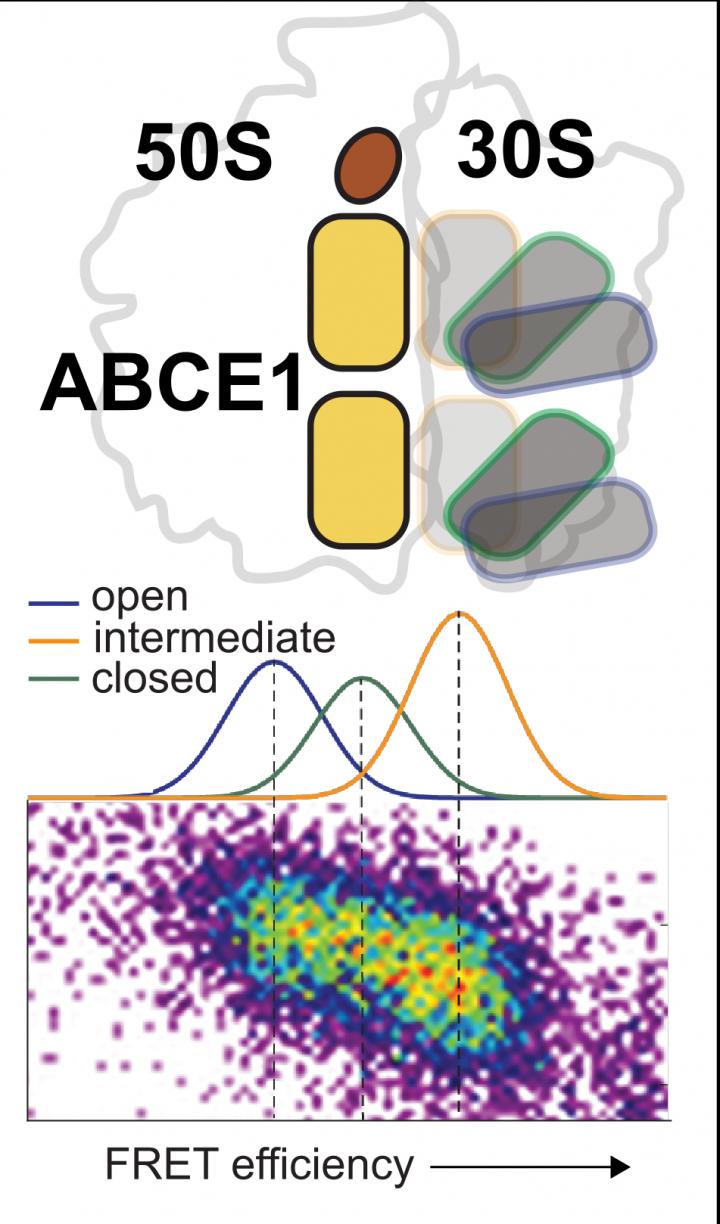
Credit: T. Cordes, LMU Munich
Ribosomes are molecular machines that produce proteins in cells. Having finished the job, the ribosomes need regenerating. This process is important for the quality of the proteins produced and thus for the whole cell homeostasis as well as for developmental and biological processes. Biochemists from Goethe University Frankfurt together with biophysicists at LMU Munich have now watched one of the most important enzymes for ribosome recycling at work – ABCE1 – and shown that it is unexpectedly versatile in terms of structure.
Ribosomes decode the genetic information from the messenger RNAs and translate it into proteins. Once they have produced a protein, but also if defective proteins have come to a halt in the ribosome, the ribosomes have to be “recycled” so that they are in good working order for a new round of synthesis. In all organisms (except bacteria), the enzyme ABCE1 coordinates this process, in which the ribosome is split into its two subunits. Biochemist Robert Tampé and LMU biophysicist Thorben Cordes, in collaboration with researchers at the University of Groningen (Netherlands), have shown that ABCE1 adopts three structural conformations to boost recycling. Their results are presented in the current issue of the journal Cell Report.
The ABCE1 enzyme can split ATP, the energy currency of cells, and use the energy released to separate the two ribosomal subunits. “Recent structural and functional data have shown that a conformational change of the enzyme, that is, a change in its spatial structure, is essential within this process for the diverse functions of ABCE1,” says Cordes. Using an integrated test approach – among others with the help of what is known as the single-molecule FRET method – his team has now observed at first hand the structural variability of ABCE1 at the level of single molecules.
In the course of this work, the researchers established that the two ATP binding sites of ABCE1 can adopt three conformations – open, intermediate and closed – which are in a state of dynamic equilibrium. Interaction of ABCE1 with both the ribosome and the ATP influences the structural dynamics of the two ATP binding sites. This results in a complex network of different states, in which ribosome and ATP shift the equilibrium in the direction of the closed conformation.
“We assume that the conformations perform functionally different roles in the dissociation of the ribosome as well as for the many other diverse functions of ABCE1,” says Cordes. “Ribosome recycling is governed by an extraordinarily complex and conserved machinery, which has medical significance as yet unimagined,” adds Robert Tampé.
###
Media Contact
Robert Tampe
[email protected]
Related Journal Article
http://dx.