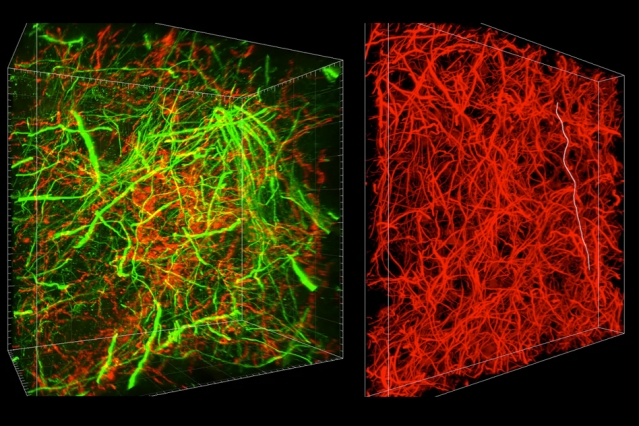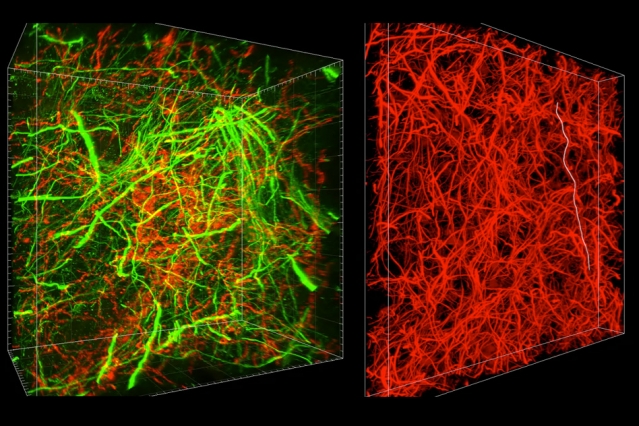
MIT researchers have developed a new technique for imaging brain tissue at multiple scales, allowing them to peer at molecules within cells or take a wider view of the long-range connections between neurons.
This technique, known as magnified analysis of proteome (MAP), should help scientists in their ongoing efforts to chart the connectivity and functions of neurons in the human brain, says Kwanghun Chung, the Samuel A. Goldblith Assistant Professor in the Department of Chemical Engineering, and a member of MIT’s Institute for Medical Engineering and Science (IMES) and Picower Institute for Learning and Memory.
“We use a chemical process to make the whole brain size-adjustable, while preserving pretty much everything. We preserve the proteome (the collection of proteins found in a biological sample), we preserve nanoscopic details, and we also preserve brain-wide connectivity,” says Chung, the senior author of a paper describing the method in the July 25 issue of Nature Biotechnology.
The researchers also showed that the technique is applicable to other organs such as the heart, lungs, liver, and kidneys.
The paper’s lead authors are postdoc Taeyun Ku, graduate student Justin Swaney, and visiting scholar Jeong-Yoon Park.
[youtube https://www.youtube.com/watch?v=9ULPT4vYOlg?rel=0&w=560&h=315]
MIT researchers have developed a new technique for imaging brain tissue at multiple scales, allowing them to peer at molecules within cells or take a wider view of the long-range connections between neurons.
Video: Melanie Gonick/MIT
Multiscale imaging
The new MAP technique builds on a tissue transformation method known as CLARITY, which Chung developed as a postdoc at Stanford University. CLARITY preserves cells and molecules in brain tissue and makes them transparent so the molecules inside the cell can be imaged in 3-D. In the new study, Chung sought a way to image the brain at multiple scales, within the same tissue sample.
“There is no effective technology that allows you to obtain this multilevel detail, from brain region connectivity all the way down to subcellular details, plus molecular information,” he says.
To achieve that, the researchers developed a method to reversibly expand tissue samples in a way that preserves nearly all of the proteins within the cells. Those proteins can then be labeled with fluorescent molecules and imaged.
The technique relies on flooding the brain tissue with acrylamide polymers, which can form a dense gel. In this case, the gel is 10 times denser than the one used for the CLARITY technique, which gives the sample much more stability. This stability allows the researchers to denature and dissociate the proteins inside the cells without destroying the structural integrity of the tissue sample.
Before denaturing the proteins, the researchers attach them to the gel using formaldehyde, as Chung did in the CLARITY method. Once the proteins are attached and denatured, the gel expands the tissue sample to four or five times its original size.
“It is reversible and you can do it many times,” Chung says. “You can then use off-the-shelf molecular markers like antibodies to label and visualize the distribution of all these preserved biomolecules.”
There are hundreds of thousands of commercially available antibodies that can be used to fluorescently tag specific proteins. In this study, the researchers imaged neuronal structures such as axons and synapses by labeling proteins found in those structures, and they also labeled proteins that allow them to distinguish neurons from glial cells.
“We can use these antibodies to visualize any target structures or molecules,” Chung says. “We can visualize different neuron types and their projections to see their connectivity. We can also visualize signaling molecules or functionally important proteins.”
High resolution
Once the tissue is expanded, the researchers can use any of several common microscopes to obtain images with a resolution as high as 60 nanometers — much better than the usual 200 to 250-nanometer limit of light microscopes, which are constrained by the wavelength of visible light. The researchers also demonstrated that this approach works with relatively large tissue samples, up to 2 millimeters thick.
“This is, as far as I know, the first demonstration of super-resolution proteomic imaging of millimeter-scale samples,” Chung says.
“This is an exciting advance for brain mapping, a technique that reveals the molecular and connectional architecture of the brain with unprecedented detail,” says Sebastian Seung, a professor of computer science at the Princeton Neuroscience Institute, who was not involved in the research.
Currently, efforts to map the connections of the human brain rely on electron microscopy, but Chung and colleagues demonstrated that the higher-resolution MAP imaging technique can trace those connections more accurately.
Chung’s lab is now working on speeding up the imaging and the image processing, which is challenging because there is so much data generated from imaging the expanded tissue samples.
“It’s already easier than other techniques because the process is really simple and you can use off-the-shelf molecular markers, but we are trying to make it even simpler,” Chung says.
Story Source:
The above post is reprinted from materials provided by MIT NEWS
The post Imaging the brain at multiple size scales appeared first on Scienmag.