In a groundbreaking study that promises to reshape our understanding of cellular sulfation processes, researchers have identified MESH1 as a pivotal enzyme responsible for the hydrolysis of the universal sulfate donor, 3′-phosphoadenosine-5′-phosphosulfate (PAPS). This discovery, anchored by meticulous biochemical experiments and high-resolution crystallographic analysis, finally uncovers the long-sought phosphatase that modulates PAPS levels, illuminating a crucial regulatory mechanism in metazoan sulfation biology.
Sulfation reactions permeate diverse biological landscapes, serving as essential post-translational modifications that govern the function of a plethora of biomolecules ranging from proteins to glycosaminoglycans (GAGs). These sulfation cascades hinge upon the availability of PAPS, a molecule whose biosynthesis has been thoroughly charted over past decades. However, the enzyme responsible for PAPS degradation remained elusive until now, presenting a critical blind spot in comprehending how cells fine-tune sulfation dynamics.
Through an integrative approach combining enzymology and structural biology, Lin and colleagues identified MESH1—a metazoan enzyme previously unassociated with sulfation metabolism—as a bona fide PAPS phosphatase. Their biochemical assays demonstrated that MESH1 catalyzes the hydrolysis of PAPS into adenosine-5′-phosphosulfate (APS) and inorganic phosphate, effectively controlling PAPS turnover and, by extension, the extent of sulfation reactions within the cell.
Crystallographic data provided a molecular snapshot of MESH1 bound to PAPS, confirming substrate specificity and revealing the intricate binding interactions that confer exquisite enzymatic activity. The structure of the MESH1-PAPS complex sheds light on the enzyme’s catalytic mechanism, highlighting key residues that orchestrate phosphate hydrolysis. This structural insight adds a vital dimension to understanding how MESH1 operates at the atomic level to govern sulfation biochemistry.
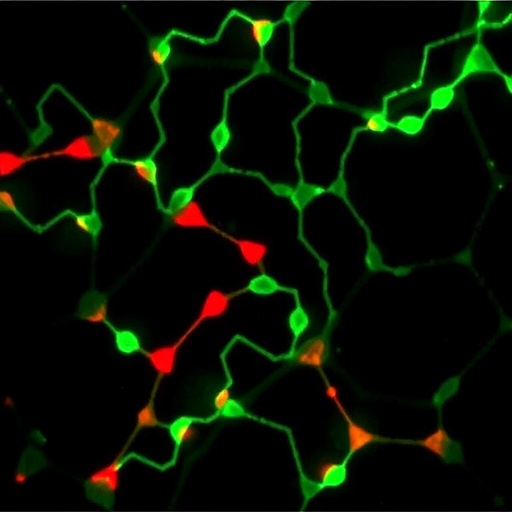
Intriguingly, subcellular localization studies demonstrated that MESH1 predominantly resides in the Golgi apparatus, the central hub for sulfotransferase activity and glycosaminoglycan modification. This spatial correlation places MESH1 strategically at the heart of sulfation machinery, poised to regulate the sulfotransferases’ substrate pool by modulating PAPS availability directly at its biosynthetic and consumption sites within the cell.
Functional experiments using a chondrogenic cell line revealed that RNA interference-mediated knockdown of MESH1 leads to a pronounced increase in sulfated glycosaminoglycan (sGAG) production. This finding underscores the enzyme’s role as a negative regulator of sulfation flux, suggesting that MESH1 activity restrains excess sGAG synthesis under physiological conditions, thus maintaining homeostatic balance.
Expanding the relevance of their findings to in vivo models, the researchers investigated the impact of Mesh1 deletion in brachymorphic mice—a genetic model exhibiting sulfation deficiencies and cartilage abnormalities. Remarkably, Mesh1 knockout animals manifested significantly elevated sGAG levels in joint cartilage alongside improved bone density, indicating that ameliorating MESH1’s inhibitory effect on sulfation positively influences musculoskeletal integrity and opens avenues for therapeutic exploitation.
The study further leveraged the model organism Caenorhabditis elegans to probe evolutionary conservation and pathological implications. In worms deficient in the 3′-phosphoadenosine 5′-phosphatase BPNT-1, which causes accumulation of cytotoxic 3′-phosphoadenosine 5′-phosphate (PAP), overexpression of MESH1 alleviated neurotoxic PAP buildup by reducing upstream PAPS quantities. This genetic interplay reveals a delicate metabolic balance maintained by phosphatases in the sulfation pathway and positions MESH1 as a critical moderator of nucleotide-sulfate metabolism.
Beyond its foundational biochemical role, MESH1’s function as a PAPS phosphatase suggests broader implications in pathophysiological contexts where sulfation dysregulation underlies disease states. Disorders marked by sulfation insufficiency—ranging from skeletal malformations to neurodegenerative conditions—may benefit from strategies targeting MESH1 to recalibrate sulfation potential and restore physiological function.
The research by Lin et al. not only fills a longstanding gap in sulfation biology but also highlights the therapeutic promise of modulating PAPS metabolism. The emerging picture is one of sophisticated cellular control, where enzymes such as MESH1 act as molecular rheostats fine-tuning sulfotransferase substrate availability to orchestrate precise sulfation patterns essential for normal development and tissue maintenance.
Moreover, the discovery opens exciting opportunities for drug discovery, as selective inhibitors or activators of MESH1 could be leveraged to modulate sulfation levels in disease-specific contexts, offering a novel class of molecular interventions. The localization of MESH1 in the Golgi further suggests that targeting intracellular compartments could yield refined control over sulfation dynamics.
This revelation also invites renewed scrutiny into related enzymes and cofactors within the sulfur assimilation and nucleotide metabolism networks, fostering a more integrated understanding of how cells manage and recycle sulfate groups. The interplay between MESH1, BPNT-1, and other metabolic nodes thus emerges as a rich area for further exploration in cell biology and medicine.
Altogether, these discoveries cement MESH1’s critical role as the long-sought PAPS phosphatase in metazoans and establish new paradigms in the regulation of sulfation. As research advances, the detailed molecular insights gained from structural and functional characterization of MESH1 promise to inspire therapeutic innovations targeting sulfation-related diseases—a testament to the enduring power of chemical biology and structural enzymology.
In light of this study, future research will likely delve into the regulatory cues governing MESH1 expression and activity, its interaction partners, and the mechanistic nuances dictating substrate selectivity. Decoding these layers of control will be vital for harnessing MESH1’s full potential in clinical settings, providing sophisticated tools to correct sulfation imbalances.
Furthermore, the translational implications for bone and cartilage disorders, neurodegeneration, and other complex syndromes linked to abnormal sulfation propel MESH1 into the spotlight as a promising molecular target. As preclinical models and eventually human studies ensue, the approach of modulating PAPS phosphatase activity could become a cornerstone of precision medicine strategies.
This landmark work serves as a clarion call to the biochemical and medical communities to rethink sulfation homeostasis in light of MESH1’s newly revealed enzymatic function, fostering cross-disciplinary collaborations that span enzymology, structural biology, and therapeutic development.
Subject of Research: Metazoan regulation of sulfation through identification and characterization of MESH1 as a PAPS phosphatase.
Article Title: MESH1 functions as a metazoan PAPS phosphatase to regulate sulfation.
Article References:
Lin, CC., Rose, J., Zhang, A. et al. MESH1 functions as a metazoan PAPS phosphatase to regulate sulfation. Nat Chem Biol (2026). https://doi.org/10.1038/s41589-026-02190-5
Image Credits: AI Generated
DOI: https://doi.org/10.1038/s41589-026-02190-5
Tags: 3′-phosphoadenosine-5′-phosphosulfate metabolismbiochemical characterization of MESH1cellular sulfation regulationcrystallographic analysis of MESH1MESH1 enzyme functionmetazoan sulfation biologyPAPS hydrolysis mechanismPAPS phosphatase activityregulation of biomolecular sulfationstructural biology of sulfation enzymessulfate donor turnover in cellssulfation post-translational modifications