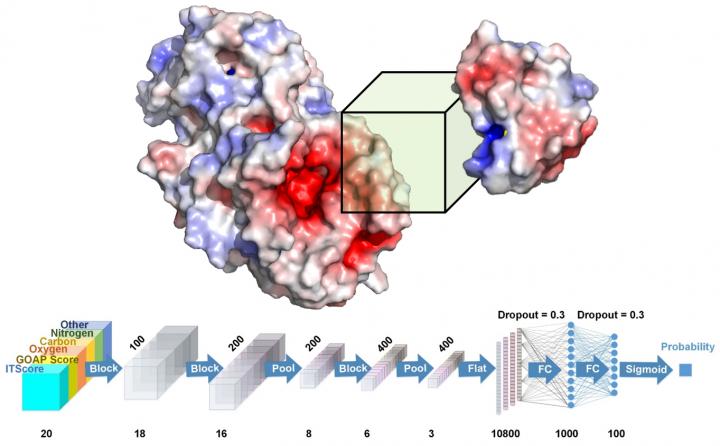
Credit: Daisuke Kihara/Purdue University
WEST LAFAYETTE, Ind. – Proteins are often called the working molecules of the human body. A typical body has more than 20,000 different types of proteins, each of which are involved in many functions essential to human life.
Now, Purdue University researchers have designed a novel approach to use deep learning to better understand how proteins interact in the body – paving the way to producing accurate structure models of protein interactions involved in various diseases and to design better drugs that specifically target protein interactions. The work is released online in Bioinformatics.
“To understand molecular mechanisms of functions of protein complexes, biologists have been using experimental methods such as X-rays and microscopes, but they are time- and resource-intensive efforts,” said Daisuke Kihara, a professor of biological sciences and computer science in Purdue’s College of Science, who leads the research team. “Bioinformatics researchers in our lab and other institutions have been developing computational methods for modeling protein complexes. One big challenge is that a computational method usually generates thousands of models, and choosing the correct one or ranking the models can be difficult.”
Kihara and his team developed a system called DOVE, DOcking decoy selection with Voxel-based deep neural nEtwork, which applies deep learning principles to virtual models of protein interactions. DOVE scans the protein-protein interface of a model and then uses deep learning model principles to distinguish and capture structural features of correct and incorrect models.
“Our work represents a major advancement in the field of bioinformatics,” said Xiao Wang, a graduate student and member of the research team. “This may be the first time researchers have successfully used deep learning and 3D features to quickly understand the effectiveness of certain protein models. Then, this information can be used in the creation of targeted drugs to block certain protein-protein interactions.”
###
Kihara has worked with the Purdue Research Foundation Office of Technology Commercialization on some of his research and technology.
About Purdue Research Foundation Office of Technology Commercialization
The Purdue Research Foundation Office of Technology Commercialization operates one of the most comprehensive technology transfer programs among leading research universities in the U.S. Services provided by this office support the economic development initiatives of Purdue University and benefit the university’s academic activities through commercializing, licensing and protecting Purdue intellectual property. The office is managed by the Purdue Research Foundation, which received the 2019 Innovation and Economic Prosperity Universities Award for Place from the Association of Public and Land-grant Universities. For more information on licensing a Purdue innovation, contact the Office of Technology Commercialization at [email protected]. The Purdue Research Foundation is a private, nonprofit foundation created to advance the mission of Purdue University.
Writer: Chris Adam, 765-588-3341, [email protected]
Source: Daisuke Kihara, [email protected]
Media Contact
Chris Adam
[email protected]
Original Source
https:/
Related Journal Article
http://dx.




