In a study published in Nature Biotechnology on Aug. 28, GAO Caixia’s group at the Institute of Genetics and Developmental Biology (IGDB) of the Chinese Academy of Sciences and her collaborators at Qi Biodesign have reported a modular, CRISPR-free base editing system, which they call CyDENT, to achieve effective base editing in the nucleus, mitochondria and chloroplast genomes of plant and human cells.
Credit: IGDB
In a study published in Nature Biotechnology on Aug. 28, GAO Caixia’s group at the Institute of Genetics and Developmental Biology (IGDB) of the Chinese Academy of Sciences and her collaborators at Qi Biodesign have reported a modular, CRISPR-free base editing system, which they call CyDENT, to achieve effective base editing in the nucleus, mitochondria and chloroplast genomes of plant and human cells.
Genome editing enables the efficient and precise modification of genetic information in living organisms, revolutionizing life science research as a whole. In recent years, the advent of base editing technology has made genome editing even more precise and predictable.
David Liu’s lab at the Broad Institute and Harvard University has pioneered the development of base editing. By fusing a nickase Cas9 with ssDNA-specific deaminases, they have successively developed the cytosine base editor (CBE) and adenine base editor (ABE) systems, which allow efficient C·G-to-T·A or A·T-to-G·C base conversions in the nuclear genome, respectively. Subsequently, they used a newly identified dsDNA-specific deaminase, DddAtox, to develop DdCBE, which enables efficient C·G-to-T·A in organellar genomes. Jin-Soo Kim’s lab built on DdCBE to develop TALED to achieve A·T-to-G·C base editing in mitochondrial genomes.
However, due to the dsDNA deamination properties of DddAtox, the DdCBE and TALED systems generate additional unintended off-target edits. Recently, WEI Wensheng’s team at Peking University has developed DddAtox-independent base editors, called mitoBEs, which improve the precision of mitochondrial base editing.
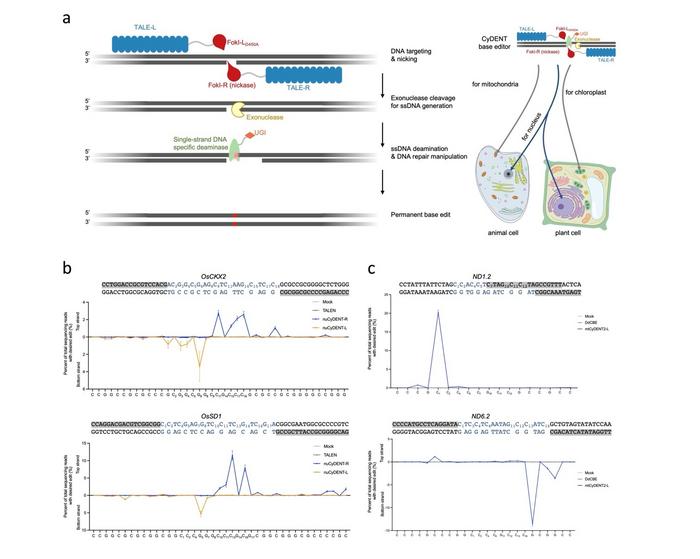
In this study, according to GAO, CyDENT consists of a pair of TALEs fused to a FokI nickase, a single-strand specific cytidine deaminase, an exonuclease and a uracil glycosylase inhibitor peptide. After the nickase nicks a TALE-guided strand of target DNA, the exonuclease next recognizes the nicked region and digests the nicked DNA strand, thereby exposing a short ssDNA fragment that serves as a substrate for single-strand DNA-specific deamination.
This strategy allows editing of single-strand DNA in the absence of a Cas9-guided R-loop structure, so base editing can be achieved without dsDNA deaminases. Since the entire CyDENT complex is RNA-free, this genome editing system enables strand-specific base editing in the nuclear and organellar genomes.
The researchers evaluated CyDENT base editing in nuclear and chloroplast target sites of rice protoplasts, and in mitochondrial sites of HEK293T cells. Effective base editing was demonstrated with editing efficiencies up to nearly 40% in the mitochondrial genome with high strand specificity. Using newly discovered ssDNA deaminases developed by the same groups, editing at TC or GC motifs was achieved, demonstrating the flexibility and modularity of the CyDENT systems.
Genome editing technologies have already made great strides in a wide range of research areas, including in the treatment of genetic diseases, the breeding of new and sustainable agricultural varieties, and the engineering of various microbiomes.
The development of CyDENT base editors expands the suite of precision genome editing technologies for both nuclear and organelle genome editing, thus facilitating better genetic designs.
Journal
Nature Biotechnology
DOI
10.1038/s41587-023-01910-9
Method of Research
Experimental study
Subject of Research
Not applicable
Article Title
Strand-preferred base editing of organellar and nuclear genomes using CyDENT
Article Publication Date
28-Aug-2023