The orchestration of cell fate decisions during the earliest stages of embryonic development is a process of remarkable complexity and precision, governed by a myriad of epigenetic mechanisms. Among these, histone modifications hold a pivotal role in fine-tuning gene expression programs that ensure faithful lineage commitment. One particular histone mark, trimethylation of lysine 27 on histone H3 (H3K27me3), has emerged as a central repressive epigenetic modification crucial for the silencing of developmental genes until their activation is precisely timed. Despite its known significance, the molecular intricacies through which H3K27me3 influences the earliest cell fate transitions remain insufficiently delineated, leaving a critical knowledge gap in developmental biology.
In pursuit of elucidating these mechanisms, researchers from the Chinese Academy of Medical Sciences (CAMS) in conjunction with the Guangzhou Institutes of Biomedicine and Health (GIBH) generated knockout models targeting Pcgf1, a gene encoding PCGF1, a core component of the non-canonical Polycomb repressive complex 1 (ncPRC1). This complex is integral to catalyzing H2AK119 mono-ubiquitination (H2AK119ub), a chromatin modification that facilitates subsequent recruitment of the Polycomb repressive complex 2 (PRC2) and its associated histone mark H3K27me3 via regulatory proteins such as JARID2. The ablation of PCGF1 in mice resulted in embryonic lethality, with no viable Pcgf1-null pups detected at birth. Investigations revealed that mutant embryos were resorbed starting at embryonic day 9.5 (E9.5), underscoring an indispensable role for PCGF1 in early development.
Intriguingly, initial developmental stages appeared morphologically and transcriptomically unperturbed at E6.5, suggesting early embryogenesis proceeds normally until a critical transition point is reached. The researchers pinpointed this juncture to gastrulation, a formative event in embryonic patterning whereby pluripotent epiblast cells commit to defined germ layers. PCGF1 deficiency caused a complete arrest of gastrulation, implicating a failure in the transition from pluripotency to lineage commitment. This suggests that in the absence of PCGF1, cells become locked within a pluripotent state, unable to activate the genetic programs necessary for differentiation.
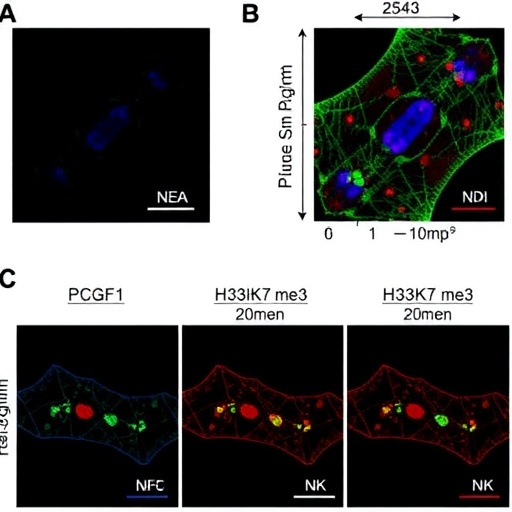
To unravel the mechanistic underpinnings of this blockade, the team employed an in vitro model of definitive endoderm (DE) differentiation derived from embryonic stem cells (ESCs). Depletion of PCGF1 in this system led to a marked reduction in H3K27me3 deposition at crucial developmental gene loci. Paradoxically, these genes exhibited only modest upregulation despite the loss of repressive marks, indicating that mere derepression was insufficient to drive lineage specification. Instead, ESCs deficient in PCGF1 retained stem cell-like characteristics, exemplified by failure to robustly activate endodermal signatures and continued high expression levels of pluripotency genes. Transcriptomic profiling during DE differentiation confirmed substantially diminished expression of endodermal markers coupled with sustained high transcript levels of core pluripotency factors, mirroring in vivo embryonic defects.
This observation prompted the hypothesis that aberrant silencing of pluripotency genes could be the critical impediment to lineage progression. Normally, pluripotency genes acquire H3K27me3 during differentiation, effectuating their transcriptional repression, which is essential to liberate cells from the undifferentiated state and permit activation of lineage-specific programs. Chromatin immunoprecipitation (ChIP) assays verified this model by demonstrating robust H3K27me3 enrichment on pluripotency gene promoters during DE differentiation in wild-type ESCs. In stark contrast, PCGF1 depletion abolished this enrichment, resulting in persistent transcriptional activity of pluripotency networks.
Further epigenomic analyses revealed that loss of PCGF1 disrupts not only the deposition of repressive H3K27me3 but also induces compensatory changes in other histone modifications. Notably, the histone methyltransferase MLL2, responsible for tri-methylating lysine 4 on histone H3 (H3K4me3)—a mark associated with gene activation—showed increased binding to chromatin in PCGF1-null ESCs. Concurrently, UTX, a histone demethylase that removes H3K27me3, was recruited at elevated levels, collectively skewing the epigenetic landscape towards one favoring pluripotency gene activation. This dysregulation of bivalent chromatin signatures disrupts the delicate balance between gene repression and activation necessary for proper embryonic development.
Taken together, these findings illuminate a sophisticated regulatory network in which dynamic modulation of H3K27me3 acts as a molecular switch essential for initiating lineage commitment. H3K27me3 silences pluripotency genes, releasing cells from an undifferentiated state, while simultaneously poised developmental genes remain bivalently marked, ready for activation upon appropriate cues. PCGF1 emerges as a gatekeeper within this paradigm, orchestrating the ncPRC1-mediated ubiquitination of H2AK119 and its downstream recruitment of PRC2, thereby ensuring the homeostasis of H3K27me3. By also constraining excessive MLL2 and UTX activity, PCGF1 preserves the epigenetic equilibrium that underlies precise gene expression dynamics during early embryogenesis.
The implications of this work extend significantly beyond developmental biology, offering valuable insights pertinent to the pathogenesis of a spectrum of developmental disorders characterized by epigenetic anomalies. Aberrant regulation of Polycomb complexes and histone modifications is increasingly linked to congenital defects and diseases, highlighting the translational importance of delineating these pathways. Moreover, this knowledge furnishes a foundation upon which strategies in regenerative medicine can be refined. Manipulating the epigenetic circuitry governing pluripotency and differentiation through modulation of H3K27me3 dynamics holds promise for enhancing the efficiency and fidelity of cell-based therapies.
In sum, the study spearheaded by CAMS and GIBH uncovers the essential function of PCGF1 and the ncPRC1 complex in safeguarding developmental progression through epigenetic regulation. It establishes a mechanistic framework whereby orchestrated chromatin remodeling balances gene repression and activation, thereby catalyzing embryonic lineage specification. These insights significantly advance our understanding of the molecular logic governing early development and set the stage for future investigations into therapeutic manipulation of epigenetic states.
Subject of Research: Epigenetic regulation of early embryonic development; role of PCGF1 and H3K27me3 in cell fate decisions.
Article Title: PCGF1 safeguards early lineage specification by coordinating chromatin regulators to balance pluripotency repression and developmental gene activation.
Web References: DOI link
Image Credits: Pan Chen
Keywords: Epigenetics, Polycomb repressive complexes, PCGF1, H3K27me3, embryonic stem cells, pluripotency, lineage commitment, gastrulation, chromatin remodeling, MLL2, UTX, developmental biology, regenerative medicine