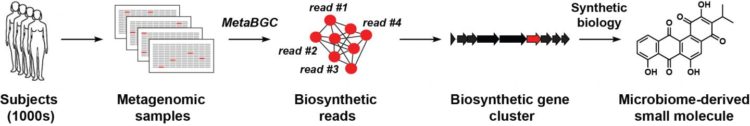
Using a massive dataset drawn from the genomes of hundreds of people, a team of Princeton researchers identified two microbes that function as powerful antibiotics.
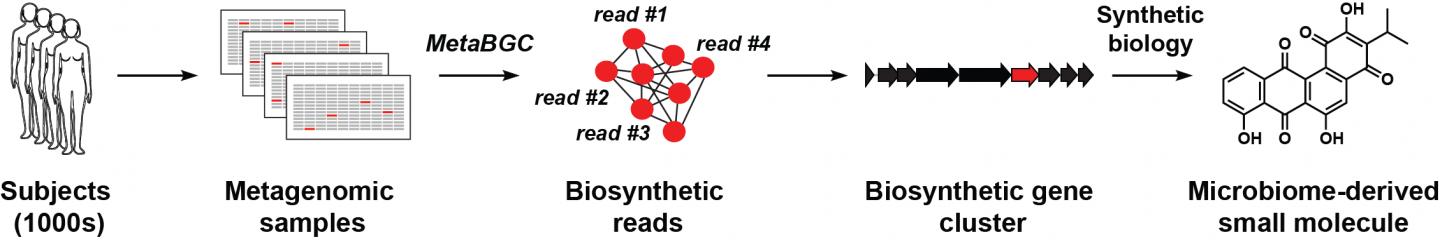
Credit: Reprinted with permission from Sakimoto, Camacho et al., Science (2019)
The microbes populating the human body play an important role in health and disease, but with few exceptions, how individual microbial species affect health and disease states remains poorly understood. A new study by Princeton researcher Mohamed Abou Donia and his colleagues, appearing in the Dec. 13 issue of the journal Science, gives scientists new tools to explore and understand the human microbiome.
Using a massive dataset drawn from the genomes of hundreds of people, the researchers identified two microbes that function as powerful antibiotics, said Donia, an assistant professor in the Department of Molecular Biology, “as potent as their clinically used relatives against neighboring microbes in the oral microbiome.” Finding novel antibiotics is important because pathogens are evolving resistance to antibiotics currently in clinical use.
The human microbiome — the identity and balance of bacterial species on human skin and mucosal surfaces — influences a variety of disease conditions, from digestive ailments to halitosis, bacterial vaginosis and eczema. The microbiome also aids immune development and the fight against pathogens. However, the human microbiome is incredibly diverse; the communities of bacteria, viruses, fungi and other tiny organisms differ according to the tissue where they live, and across human populations and individuals. It’s unclear what constitutes a normal, healthy microbiome, much less how one might go about bringing a sick one back into balance.
A common approach to solving this problem is to culture an individual microbe in the lab and explore how it contributes to health or disease states. Unfortunately, it can be difficult to identify and isolate very rare species, or find the conditions necessary to support their growth outside their natural niche. To do this with every species would be a daunting task. Alternatively, scientists can examine the microbiome in situ, with the aim of describing its individual components and how they interact.
One way microbes communicate — and do battle — with each other and with human cells is through biologically active small molecules.
“Our long-term goal is to define the chemical space of the human microbiome,” explained Donia. His group set out to identify the set of genes that manufacture such chemicals (termed a biosynthetic gene cluster, or BGC) directly in clinical samples. This would allow scientists to listen in on the chemical conversation taking place, and discover who is speaking and when.
Led by co-first authors Yuki Sugimoto, a postdoctoral research associate, and graduate student Francine Camacho, the researchers developed computer algorithms that can detect BGCs by analyzing and interpreting data sets known as “metagenomic sequencing data,” genetic sequences obtained from the tissues or excretions of hundreds of human subjects. Some metagenomic data sets are drawn from clinical samples taken from diverse populations, including persons in different states of health or disease, or people in different geographical locales. Intensive analysis is needed to make sense of the rich but often fragmentary information contained in these data sets.
Donia’s team began by identifying genes essential for the synthesis of a particular molecule or chemical of interest, then using computational algorithms to sort through metagenomic data for similar (homologous) genetic sequences, and grouping these sequence fragments together. They then assessed the prevalence of each group in the human population, and used the grouped sequences to piece together full-length BGCs. Importantly, this approach allowed identification of novel BGCs even if they are extremely rare.
To validate this approach, the researchers investigated whether they could detect BGCs involved in the synthesis of type II polyketides. This class of chemicals, which includes the anti-cancer drug doxorubicin and several antibiotic drugs, was previously found in soil bacteria but had never before been found in bacteria of the human microbiome.
“To our surprise, we discovered 13 such gene clusters, which are widely distributed in the gut, oral and skin microbiome of people all the way from the U.S. to Fiji,” said Donia. To test if these newly identified BGCs actually make type II polyketides, the researchers selected two of the BGCs and inserted their genes into bacteria that can be easily grown in the lab, then used mass spectrometry to detect any new chemical products. These compounds were then purified and tested for antibiotic or anticancer activity.
“Two of the five new molecules we discovered are potent antibiotics,” said Donia, “revealing a potential mechanism for niche competition and defense against intruders and pathogens.” More work will be needed to discover the biological activity of the other three molecules, and the role of all five in human health or disease. Such studies may uncover new pathways for interaction between microbes, or between the microbiome and its human host.
With this technology, it is now possible to mine our own microbiomes for drug discovery or novel biological interactions. What other treasures might this type of analysis reveal? As Donia observed, “This was only one clinically important class of molecules that we went after — there are dozens more to do, and we can’t even start to predict what we will discover!”
###
“A metagenomic strategy for harnessing the chemical repertoire of the human microbiome,” by Yuki Sugimoto, Francine R. Camacho, Shuo Wang, Pranatchareeya Chankhamjon, Arman Odabas ’17, Abhishek Biswas, Philip D. Jeffrey and Mohamed S. Donia, appears in the Dec. 13 issue of the journal Science and was released online Oct. 3 (DOI: 10.1126/science.aax9176). This work was supported the National Institutes of Health Director’s New Innovator Award (1DP2AI124441), the Pew Biomedical Scholars Program, and a Focused Research Team on Precision Antibiotics Award by the School of Engineering and Applied Science at Princeton University. The researchers are all at Princeton University, in the Department of Molecular Biology, the Lewis-Sigler Institute for Integrative Genomics, the Department of Chemical and Biological Engineering, or Princeton Research Computing.
Media Contact
Liz Fuller-Wright
[email protected]
609-258-5729
Related Journal Article
http://dx.