DURHAM, N.C. — Fungal infections can be devastating to human health, killing approximately 150 people every hour, resulting in over a million deaths every year, more than malaria and tuberculosis combined.
Unfortunately the antifungal drug arsenal is limited, with many of the best drugs more than 50 years old.
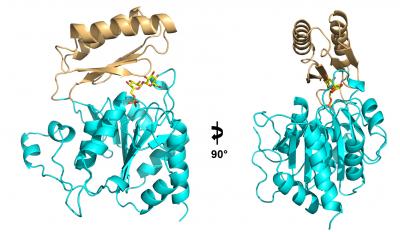
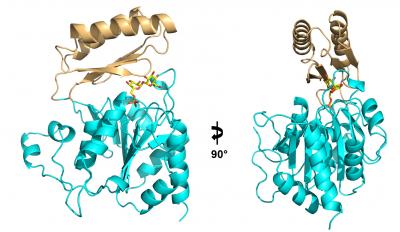
The search for new antifungals has recently alighted on a simple biological pathway, the production of trehalose, a chemical cousin to table sugar that pathogenic fungi need to survive in their human hosts. A team of Duke researchers has solved the structure of an enzyme that is required to synthesize this fungal factor.
When the enzyme is blocked, pathogenic fungi are unable to endure the stressful leap from the outside world into the human body.
The research, appearing this week in the Proceedings of the National Academy of Sciences, paves the way for designing new antifungal drugs against an enzyme that is essential to the three most deadly fungi: Cryptococcus, Candida, and Aspergillus.
"We found that the active site of this enzyme is identical in each of these pathogenic fungi, so they all use the same mechanism and have the same residues that are necessary for the production of trehalose," said senior study author Richard G. Brennan, Ph.D., professor and chair of biochemistry at Duke University School of Medicine.
"By targeting those active sites, we hope to create a drug with broad-spectrum effects," Brennan said. "If you could kill one fungus, you're going to kill them all; at least if you can get the drug inside."
The development of antifungal drugs has lagged far behind that of other antimicrobials. Because fungi are eukaryotic multicellular organisms like humans, they share a lot of the same genes, proteins, and pathways. As a result, drugs that go after a fungal protein can also harm the human, triggering devastating side effects. These drug toxicities are one of the main reasons that antifungal therapies ultimately fail, either in the clinic or earlier in development. Luckily, the enzymes for synthesizing the trehalose sugar aren't present in people, so scientists can target them with less concern for collateral damage.
Pathogenic fungi use trehalose as a kind of molecular heat shield. Organisms like Cryptococcus, Candida, and Aspergillus spend most of their time either down in the dirt or out on the laboratory bench, where the temperature is a comfortable 30 degrees Celsius. But when they move into an unlucky host, they are suddenly exposed to a steamy 37 degrees. Trehalose protects the fungi's delicate proteins and membranes from falling apart in all that heat.
Brennan and his colleagues, including John R. Perfect, M.D., Chief of the Division of Infectious Diseases at Duke University Medical Center, figured that if they could stop trehalose from being made, they could eliminate the pathogen's protection, such that every time the fungi jump into a host they are sending themselves to a fiery death.
Trehalose is a disaccharide or a double sugar, two sugar molecules that are made and linked together by two enzymes called Tps1 and Tps2, which do their jobs one after the other. Before the scientists could target either of the biosynthetic enzymes, they needed to know its structure.
Yi Miao, PhD, a graduate student in Brennan's lab, followed the instructions written in the genetic sequence of Cryptococcus, Candida, and Aspergillus to build Tps2 (the latter of the two biosynthetic enzymes) from scratch. Then he used a technique called x-ray crystallography to produce an atomic-level three-dimensional structure of the essential molecule.
Mao took several pictures of Tps2 to produce a kind of time-lapse video of the molecule in action: first, as it sat idle; next, as it bound a precursor molecule called trehalose-6-phosphate, popping off its phosphate group to create trehalose; and finally, its job done, as it released trehalose and was alone again.
Afterwards the researchers generated a series of mutations in Tps2 to see if they could impair its ability to produce trehalose, and as a result render pathogenic fungi susceptible to heat shock. They showed that a wide variety of mutations could inactivate the enzyme and create fungi unable to survive the jump from 30 to 37 degrees.
"It appears that this enzyme has become so specific that it has basically evolved itself into a corner." Brennan said. "That's good for us, because if it tries to evolve resistance to an antifungal drug, it will likely lose its activity in the process, so the drug might not be able to destroy it but it will be a lousy enzyme, and the pathogen will die anyway."
Brennan says he and his colleagues are now using their structures of Tps2 to conduct "rational" drug design to create new antifungals that can kill a broad spectrum of pathogenic fungi. They are also performing high-throughput screening for small molecules or other pharmaceuticals that can generate the same effect.
###
This research was supported by the National Institutes of Health (1P01 AI104533, R01 AI073896, and R01 AI093257), and the American Lebanese Syrian Associated Charities, St. Jude Children's Research Hospital.
CITATION: "Structures of Trehalose-6-Phosphate Phosphatase From Pathogenic Fungi Reveal the Mechanisms of Substrate Recognition and Catalysis," Yi Mao, Jennifer L. Tenor, Dena L. Toffaletti, Erica J. Washington, Jiuyu Liu, William R. Shadrick, Maria A. Schumacher, Richard E. Lee, John R. Perfect, and Richard G. Brennan. Proceedings of the National Academy of Sciences, Online June 13, 2016. DOI: 10.1073/pnas.1601774113
Media Contact
Karl Bates
[email protected]
919-681-8054
@DukeU
http://www.duke.edu