A groundbreaking study has shed new light on the astonishing diversity of protein structures and their folds in nature. Researchers set out to reveal the extent to which nature has explored the vast landscape of possible protein topologies. The results have unveiled an astounding array of unexplored protein folds, expanding our understanding and uncovering the depth of the protein universe.
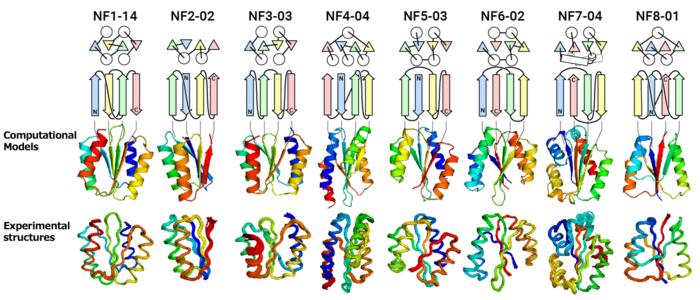
Credit: Shintaro Minami, Naohiro Kobayashi, Toshihiko Sugiki, Toshio Nagashima, Toshimichi Fujiwara, Rie Tatsumi-Koga, George Chikenji & Nobuyasu Koga, “Exploration of novel αβ-protein folds through de novo design”, Nature Structural & Molecular Biology (2023)
A groundbreaking study has shed new light on the astonishing diversity of protein structures and their folds in nature. Researchers set out to reveal the extent to which nature has explored the vast landscape of possible protein topologies. The results have unveiled an astounding array of unexplored protein folds, expanding our understanding and uncovering the depth of the protein universe.
This research has been published in the journal Nature Structural and Molecular Biology on July 3, 2023.
Proteins, the building blocks of life, fold into specific three-dimensional structures, enabling them to carry out their biological functions. The three-dimensional structures of proteins are dictated by their amino acid sequences. While experimental techniques have successfully unraveled the structures of numerous proteins over the years, the discovery of new protein folds, defined by the arrangement and connectivity of α-helices and β-strands, has become increasingly infrequent. This raises the question: how extensive is the protein fold space not explored by nature?. In attempts to address this long-standing question, theoretical studies have been conducted; however, experimental validation is lacking.
To address this question, the research team embarked on a study combining theoretical prediction for novel protein folds with experimental validation of their de novo designs.
The research team devised rules based on physical chemistry and protein structure data to theoretically predict possible protein folds. These rules were then employed to predict novel αβ-folds, which consist of a four to eight stranded β-sheet, not yet observed in the current Protein Data Bank (PDB). This led to the identification of a total of 12,356 novel folds. The team then attempted to computationally design proteins for the predicted novel folds from scratch to assess the foldability and fidelity of the novel folds.
“We attempted to computationally design proteins with all of the predicted folds that have a four-stranded β-sheet, including one forming a knot-like structure,” said Shintaro Minami, a researcher at Exploratory Research Center on Life and Living Systems (ExCELLS). “When designing proteins, we did not expect all of them, especially knot forming ones, to fold into the structures as anticipated.”
The results of experimental testing were surprising (See Figure). “For all of the folds, the computationally designed protein structures closely matched the experimental structures,” said Naohiro Kobayashi, a senior research fellow at RIKEN.
These findings suggest the existence of at least approximately 10,000 unexplored foldable αβ-folds, a significant revelation considering only 400 αβ-folds have been observed in nature. This suggests that many potential folds remain uncharted in the protein folding space.
These results have given rise to several hypotheses about the structure and evolution of proteins. One hypothesis is that proteins may have not been present in biology long enough for all possible folds to have been explored. Another hypothesis is that protein folds in nature are inherently biased due to all life on Earth having descended from a common ancestor. “Proteins may have evolved by repeatedly re-using specific folds while expressing different functions. If extraterrestrial life does exist, it might be utilizing a different set of protein folds,” said George Chikenji, an assistant professor at the Nagoya University.
Proteins are known for their diverse functions, which are generated from the diversity of protein three-dimensional structures. This study has revealed the existence of at least approximately 10,000 uncharted foldable αβ-folds in nature. “The design of proteins with these novel folds will lead to an even greater diversity of structures. This would pave the way for the de novo design of functional protein molecules, lead to breakthroughs in drug development, enzyme design, and other areas,” said Nobuyasu Koga, a professor at the Exploratory Research Center on Life and Living Systems (ExCELLS), National Institutes of Natural Sciences (NINS).
The research team includes Shintaro Minami (formerly at ExCELLS NINS); Naohiro Kobayashi from RIKEN, Toshihiko Sugiki from the Institute for Protein Research (IPR), Osaka University (currently Kitasato University), Toshio Nagashima from RIKEN, Toshimichi Fujiwara from IPR, Osaka University, Rie Tatsumi-Koga from ExCELLS NINS (currently IPR, Osaka University), George Chikenji from the Nagoya University, Nobuyasu Koga from ExCELLS NINS, Institute for Molecular Science (IMS) NINS, SOKENDAI (The Graduate University for Advanced Studies) (currently IPR, Osaka University).
For more detailed information, please refer to the published paper: “Exploration of novel αβ-protein folds through de novo design” URL: https://www.nature.com/articles/s41594-023-01029-0
Journal
Nature Structural & Molecular Biology
DOI
10.1038/s41594-023-01029-0
Method of Research
Experimental study
Subject of Research
Cells
Article Title
Exploration of novel αβ-protein folds through de novo design
Article Publication Date
3-Jul-2023