In a groundbreaking advancement for agricultural genomics, researchers have unveiled a meticulously assembled genome of JM44, a prominent Chinese bread wheat cultivar celebrated for its superior dough qualities. This new genomic resource, published in Nature Plants, transcends previous assemblies with its reference-level quality, achieving a remarkable quality value of 66.74. It provides an unparalleled, detailed representation of the notoriously complex gluten gene regions, which are integral to wheat’s end-use qualities, such as baking performance and dough elasticity. This monumental step promises to revolutionize our understanding of wheat genetics, with profound implications for global food security and wheat breeding.
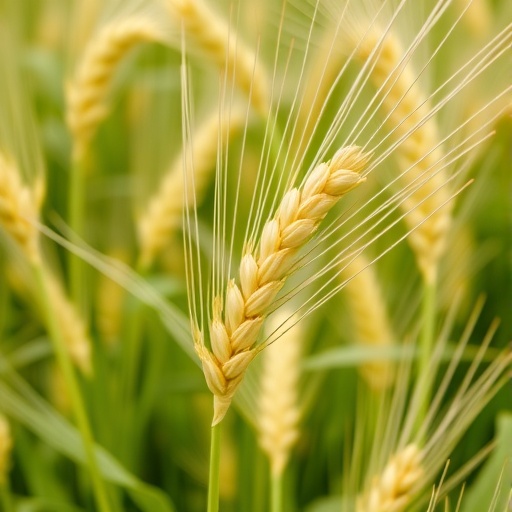
Bread wheat, with its complex hexaploid genome, has long posed formidable challenges to genome sequencing due to its massive size and abundant repetitive sequences. The JM44 cultivar is highly valued by breeders and bakers alike for its exceptional dough strength and baking performance, traits largely governed by gluten proteins encoded in the wheat genome. Gluten comprises two major protein classes, glutenins and gliadins, whose genetic architectures are complex and have evolved under both natural and human selection forces. By successfully decoding these regions with high fidelity, the JM44 assembly lays bare the intricate genetic landscape that underpins superior dough performance.
Microsynteny analysis, a comparative genomics approach assessing the conservation of gene order and orientation, was employed across the Triticum–Aegilops species complex to unravel the evolutionary dynamics of gluten genes. The study revealed that high-molecular-weight (HMW) glutenin subunit loci, pivotal for dough elasticity and strength, are extraordinarily conserved across wheat and its relatives. This conservation underscores the fundamental and perhaps indispensable role these genes play in wheat’s functional qualities. Conversely, loci encoding low-molecular-weight (LMW) glutenins and α-/β-gliadins bore the marks of extensive structural variation, suggesting these genes have been more fluid and adaptable during wheat evolution.
Such structural variation at LMW glutenin and gliadin loci reflects the intricate interplay between natural variation and human-mediated selection. The data suggest that these genomic regions have been preferentially targeted during wheat domestication and breeding, as they directly influence parameters such as extensibility and viscosity of dough. These traits are critical for a variety of wheat-based culinary applications worldwide, from bread loaves and noodles to pastries and other bakery products. Understanding the evolutionary history and selection footprints within these loci now offers breeders precise genomic targets to fine-tune wheat quality traits.
Beyond single-gene analyses, the team examined the nature of genetic interactions, or epistasis, among gluten gene loci in both traditional wheat landraces and modern cultivars. Intriguingly, epistatic effects—where the presence or expression of one gene influences another—were substantially stronger in contemporary cultivars compared to landraces. This finding implicates breeding practices over recent decades not just in selecting individual beneficial alleles but also in enhancing gene-gene interactions that synergistically improve dough quality. The results highlight epistatic selection as a critical yet often overlooked dimension of crop improvement.
The reference-level assembly of JM44 enables precise dissection of gluten gene clusters, many of which reside in highly repetitive and structurally complex genomic regions. Previous assemblies faltered in resolving these areas, leaving critical gaps in our knowledge. By harnessing advanced sequencing technologies and novel assembly algorithms, this study overcame these hurdles, producing a complete and contiguous genome assembly. Such technological strides set a new benchmark for future wheat genomics projects and facilitate genome-informed breeding strategies previously unattainable.
The study’s revelations about genome structure and evolutionary trajectories have a dual significance: first, they enhance the fundamental biological understanding of gluten gene evolution; second, they provide practical tools to breeders aiming to meet the growing demand for wheat varieties with targeted end-use qualities. The ability to manipulate specific gluten gene variants or combinations with precision promises not only improved baking characteristics but also potential nutritional enhancements, bolstering wheat’s status as a staple crop for global populations.
Moreover, the conservation of HMW glutenin subunits underscores their critical role as a genetic backbone for dough strength, suggesting that future breeding should maintain these loci intact while focusing variation efforts on more plastic regions such as LMW glutenins and gliadins. This balance between conserved and variable components could optimize wheat quality without compromising yield or adaptability. Breeders now have a clearer map of where to deploy genetic diversity to achieve desired phenotypes.
The identification of epistatic interactions as a key component of modern cultivar performance invites new frameworks for genetic selection. Traditional breeding methods often emphasize additive effects of individual genes, but this study demonstrates the necessity of considering gene networks and their combinatorial impacts on phenotype. Moving forward, integrating epistasis into predictive breeding models could substantially accelerate the development of wheat varieties tailored for specific baking qualities or environmental conditions.
In addition to direct breeding applications, the JM44 genome assembly enables comparative evolutionary studies across the Triticum genus and its relatives in the Aegilops complex. By tracing structural changes and gene duplications, researchers can infer adaptive responses to historical environmental pressures and human cultivation practices. These insights deepen our appreciation of how wheat evolved to meet diverse agronomic and culinary demands, informing not only breeding but also conservation strategies of wheat genetic resources.
As global challenges such as climate change threaten crop resilience, genetic resources like the JM44 assembly become invaluable. It provides a detailed genetic roadmap to harness both ancient variation and recent innovations in wheat for breeding climate-resilient, high-quality cultivars. The ability to combine superior end-use quality with stress tolerance traits at the genomic level offers hope for sustaining wheat productivity and food quality in a changing world.
The technical achievement of reconstructing such a large and complex polyploid genome at reference quality is a testament to advancements in sequencing technology, bioinformatics, and international collaboration. It reflects an era where crop genomes are no longer “black boxes” but accessible blueprints for precision agriculture. The JM44 genome sets a new gold standard not only for wheat but also for other cereal crops with complex genomes, potentially catalyzing a wave of similar breakthroughs.
Furthermore, the study’s findings carry implications beyond wheat. Understanding the genetic basis of protein quality and epistatic interactions offers paradigms applicable to other crop species where protein composition influences nutritional and processing qualities. This cross-disciplinary relevance enhances the study’s significance within plant genomics and crop improvement domains.
In conclusion, the high-quality JM44 bread wheat genome assembly emerges as a transformative resource, bridging fundamental genomics, evolutionary biology, and applied breeding. Its detailed portrayal of gluten gene architecture and interactions reveals the complex genetic foundation of wheat quality and exemplifies the power of genomics to unlock crop potential. This milestone paves the way for a new generation of wheat breeding strategies that can sustainably satisfy the world’s nutritional needs while honoring the rich evolutionary history of this essential staple.
Subject of Research:
Wheat genomics and the genetic basis of end-use quality traits.
Article Title:
A high-quality bread wheat genome unravels the adaptive evolution of wheat end-use quality.
Article References:
Cao, X., Zhang, J., Guo, Y. et al. A high-quality bread wheat genome unravels the adaptive evolution of wheat end-use quality. Nat. Plants (2026). https://doi.org/10.1038/s41477-026-02288-7
DOI:
https://doi.org/10.1038/s41477-026-02288-7
Tags: agricultural genomics in wheatcomplex repetitive sequences in wheatevolution of wheat end-use traitsglutenin and gliadin genetic architecturehexaploid wheat genome sequencinghigh-quality wheat genome assemblyJM44 Chinese bread wheatmicrosynteny in wheat geneticswheat breeding for baking performancewheat dough quality geneticswheat genome and food securitywheat gluten gene regions