Credit: UNIST
Membrane proteins make up approximately a quarter of all gene products and are the targets of over 50% of all modern pharmaceutical drugs. The inner mitochondrial membrane (IMM) proteome plays a central role in maintaining mitochondrial physiology and cellular metabolism. Despite their importance, there has been no method to reveal the topology of mitochondrial membrane proteins in live cells, until now.
A recent study, affiliated with UNIST has developed a new technique to understand the correct architectures of IMM proteins, using special chemical tools. By solving the most difficult stage of new drug development, their work will help speed the development of new therapeutics and cures.
This research has been led by the team of Professor Hyun-Woo Rhee of Chemistry at UNIST in collaboration with Professor Jong-Seo Kim at Center from the Center for RNA Research, within the Institute for Basic Science (IBS) at Seoul National University and Professor Jeong-Kon Seo of UNIST Central Research Facilities (UCRF). The results of the study have been appeared in the March 15th edition of the Journal of the American Chemical Society (JACS).
IMM is one of the most active sites for cellular metabolism and it is deeply related for various human metabolic diseases including cancer and neurodegenerative diseases. Therefore, it is crucial to understand the correct architecture of the IMM proteome in live cells for successful and efficient development of mitochondria-targeted therapeutics.
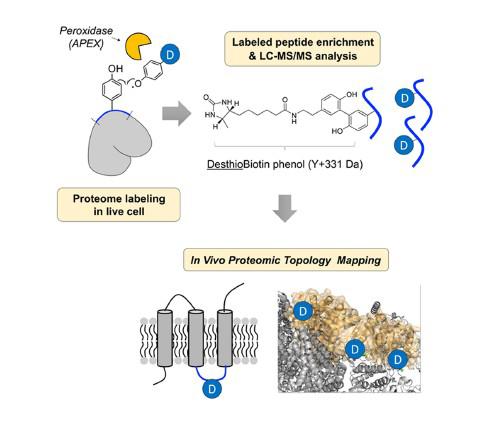
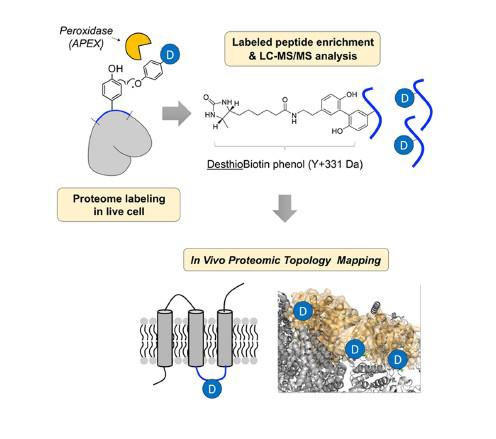
In the study, Professor Lee and his research team revealed the in vivo topological direction of 135 IMM proteins, using an in situ-generated desthiobiotin-phenoxyl radical probe with genetically targeted peroxidase (APEX).
"The determination of membrane protein structure is one of the most challenging tasks in protein structure analysis," says Professor Lee. "Our identification of structural information on the mitochondrial inner-membrane proteome can provide valuable insights for the architecture and connectome of the IMM proteome in live cells."
The research team designed a new chemical probe, desthiobiotin-phenol and applied it to the IMM proteins in live cells. Then, they identified the structure of membrane proteins via mass spectrometry (MS).
Peroxidase can make phenoxyl radical when react with hydrogen peroxide. Then, phenoxyl radical can react with tyrosine residue on the proximal protein forming covalent bond. In the study, the research team obtained the topology information by analyzing labeled tyrosine site of membrane protein.
The majority of protein sequence analysis today uses mass spectrometry (MS), which digests the protein sample into peptides using an appropriate enzyme. Previous analyses, which used genetically-targeted ascorbate peroxidase (APEX) could not resolve structural identification because these analyses performed based on the unlabeled peptide detection. However, only labeled peptide can provide useful structural information, according to the research team.
Unlike biomolecules that are labeled with biotin-phenol, proteins and other targets that are labeled with desthiobiotin-phenol can be eluted without harsh, denaturing conditions. Moreover, as the number of available membrane protein structure samples, obtained via MS increases, the efficiency of structural identification of membrane proteins also increases.
Owing to the short lifetime of phenoxyl radicals generated in situ by submitochondrial targeted APEX and the impermeability of the IMM to small molecules, the solvent-exposed tyrosine residues of both the matrix and intermembrane space (IMS) sides of IMM proteins were exclusively labeled with the radical probe in live cells by Matrix-APEX and IMS-APEX, respectively and identified by mass spectrometry.
Through this analysis, the research team confirmed 58 IMM protein topologies and determine the topological direction of 77 IMM proteins whose topology at the IMM has not been fully characterized.
###
This work has been supported by the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI) funded by the Ministry of Health & Welfare of Korea. And also supported by the Institute for Basic Science from the Ministry of Science, ICT, and Future Planning of Korea.
Journal Reference
Song-Yi Lee, et al., "Architecture Mapping of the Inner Mitochondrial Membrane Proteome by Chemical Tools in Live Cells", JACS, (2017).
Media Contact
JooHyeon Heo
[email protected]
82-522-171-223
############
Story Source: Materials provided by Scienmag