Seeing the unseen: HKU scientists develop a new chemical tool that sheds light on how proteins recognise and interact with each other
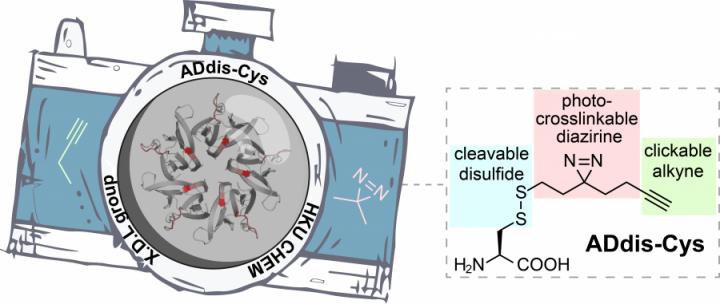
Credit: The University of Hong Kong
A research group led by Professor Xiang David LI from the Research Division for Chemistry and the Department of Chemistry, The University of Hong Kong, has developed a novel chemical tool for elucidating protein interaction networks in cells. This tool not only facilitates the identification of a protein’s interacting partners in the complex cellular context, but also simultaneously allows the ‘visualisation’ of these protein-protein interactions. The findings were recently published in the prestigious scientific journal Molecular Cell.
In the human body, proteins interact with each other to cooperatively regulate essentially every biological process ranging from gene expression and signal transduction, to immune response. As a result, dysregulated protein interactions often lead to human diseases, such as cancer and Alzheimer’s disease. In modern biology, it is important to comprehensively understand protein interaction networks, which has implications in disease diagnosis and can facilitate the development of treatments.
To dissect complex protein networks, two questions need to be answered: the ‘who’ and ‘how’ of protein binding. The ‘who’ refers to the identification of a protein’s interacting partners, whereas the ‘how’ refers to the specific ‘binding regions’ that mediate these interactions. Answering these questions is challenging, as protein interactions are often too unstable and too transient to detect. To tackle this issue, Professor Li’s group has previously developed a series of tools to ‘trap’ the protein-to-protein interactions with a chemical bond. This is possible because these tools are equipped with a special light-activated ‘camera’ – diazirine group that capture every binding partner of a protein when exposed to UV light. The interactions can then be examined and interpreted. Unfortunately, the ‘resolution’ of this ‘camera’ was relatively low, meaning key information about how proteins interact with each other was lost. To this end, Professor Li’s group has now devised a new tool (called ADdis-Cys) that has an upgraded ‘camera’ to improve the ‘resolution’. An alkyne handle installed next to the diazirine makes it possible to ‘zoom in’ to clearly see the binding regions of the proteins. Coupled with state-of-the-art mass spectrometry , ADdis-Cys is the first tool that can simultaneously identify a protein’s interacting partners and pinpoint their binding regions.
In the published paper, Professor Li’s lab was able to comprehensively identify many protein interactions — some known and some newly discovered — that are important for the regulation of essential cellular processes such as DNA replication, gene transcription and DNA damage repair. Most importantly, Professor Li’s lab was able to use ADdis-Cys to reveal the binding regions mediating these protein interactions. This tool could lead to the development of chemical modulators that regulate protein interactions for treating human diseases. As a research tool, ADdis-Cys will find far-reaching applications in many areas of study, particularly in disease diagnosis and therapy.
###
For more information about the paper “A tri-functional amino acid enables mapping of binding sites for posttranslational modification-mediated protein-protein interactions” published in Molecular Cell, please visit: https://www.cell.com/molecular-cell/fulltext/S1097-2765(21)00268-9
For more information about Professor Xiang David Li and his research group, please visit their group’s webpage: https:/
Media Contact
Casey To
[email protected]
Original Source
https:/
Related Journal Article
http://dx.